Integrated Genomics Laboratory Services Overview
We provide a coordinated pipeline of services which can be accessed at whatever point the researcher requires. Our services include in depth consultation, nucleic acid extraction, bulk and single cell sequencing, targeted gene analyses, and data processing and QC.
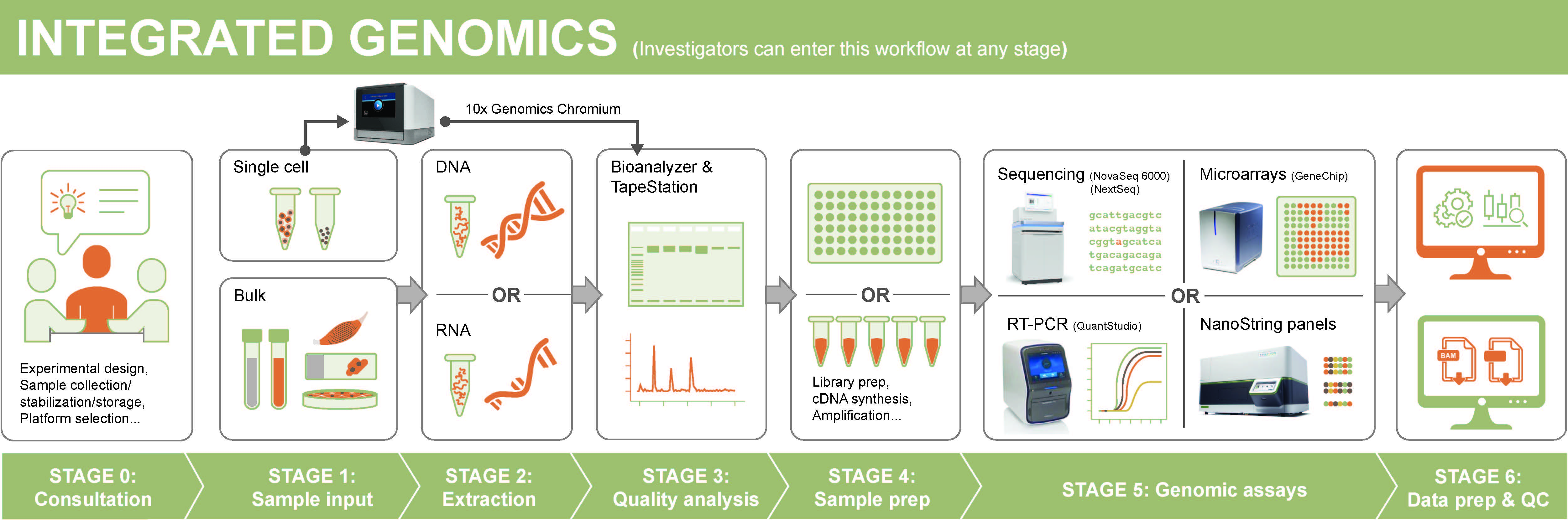
Services
-

Select image to learn more real-time PCR services 
Select image to learn more about NanoString services 
Select image to learn more about single cell and nuclei preparation services 
Select image to learn more about nucleic acid isolation services 
Select image to learn more about nucleic acid assessment services -

Select image to learn more about sequencing services 
Select image to learn more about library preparation services -

Select image to learn more about cell line authentication services 
Select image to learn more oligo ordering