MR Epilepsy WWO Neuro Protocol
Last updated: 4/15/22
Charge as: Brain WWO
Scanner preference: 3T only. If patient has an implant unsafe for 3T, OK to scan on MR2 1.5T Ingenia.
Coil: Head
-
Exam data must include: Age/DOB, Exam Date, Gender, and ID
NeuroQuant System-specific Parameter Details
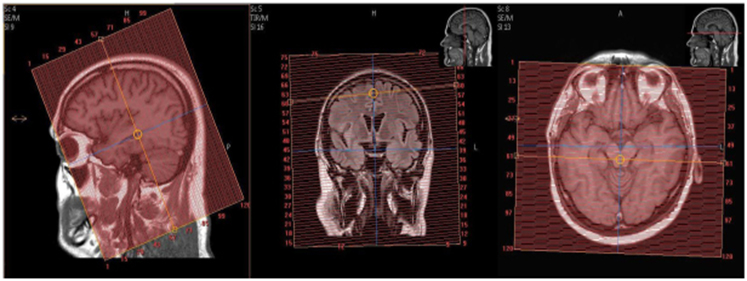
Landmark at nasion/glabella (±50mm), you must re-landmark in the brain if another body part is scanned first.
- Keep the 3D box as straight as possible.
- No patients with shunts or major artifact-causing items.
- Braces are usually okay, if there is not a great deal of motion, keep head tightly padded.
- Kyphosis prop up hips not head coil.
- Keep patient at Isocenter For patients with small heads and long necks or large heads:
- Keep FOV box positioned higher than normal but not beyond ±50mm from glabella
- May need to reduce/enlarge the FOV for the individual (not beyond 24 - 25.6)
- FOV must include all of scalp, nose and chin.
Required Scan Parameters
Following parameters remain consistent:
1. Plane – sagittal
2. Mode – 3D
3. T1 weighted - Always
4. Matrix – 192 x 192
5. NEX / NSA – 1
6. Slice thickness – 1.2 mm
7. Spacing – 1.2 mm
8. Number of slices – 160 - 170
9. FOV 24 – 25.6NOTE: Some NeuroQuant parameters vary depending on scanner manufacturer & field strength
NeuroQuant Failures (And How to Avoid Them)
Patient Motion
- Run NeuroQuant sagittal 3D first
- Pad head securely
- Use all motion reduction techniques except changing scan parameters
Artifacts
- Surgical resections, shunts, metal (some are not compatible)
- Incompatible with NeuroQuant
- Tumors > 15cc
- Contrast media - scan Non Contrast
Low SNR
- Follow recommended scan parameters
Tissue contrast Error
- Put saline bags on either side of patient's head
- Sat bands
- Follow recommended scan parameters
Incorrect landmark
- Must landmark at nasion/glabella
- Can be ± 50mm from Nasion - should be as close as possible in all 3 planes
- Re - landmark, if C-spine was done first as part of a double study
Network Issue
- Echo test failure – call your network admin
- Resend series
- Error report “Series is missing slices”
Unsupported MR Sequence
- Delete incorrect series from queue monitor
- Resend Sag 3D T1 series
-
Sending Neuroquant Images
Step 1. No Post Processing or Reformatting
Do not reformat or do any post-processing to the NeuroQuant Series. This will cause errors in the DICOM tags that can cause NeuroQuant to reject the Images.Step 2. Send Images Immediately to the Appropriate Node
As soon as you acquire the NQ series, send it to the appropriate node. This allows processing time while the study is completed. NQ processing takes about 15-30 minutes- Multiple Sclerosis: Send to NQ_MS.
- Epilepsy: Send to NQ_EPILEPSY
- Dementia: Send to NQ_DEMENTIA
Step 3. Check to make sure the correct report(s) are generated in PACS. If there are errors, email Joe and Wayne.
- NQ_MS. Generates 1 report: Multistructure Atrophy
- NQ_EPILEPSY. Generates 3 reports: Age-Related Atrophy, Multistructure Atrophy, and Triage Brain Atrophy
- NQ_DEMENTIA. Generates 3 reports: Hippocampal Asymmetry, Multistructure Atrophy, and Triage Brain Atrophy
| ?Plane | Weighting | Mode | Slice (mm) | Gap (mm) | FAT SAT | FOV (cm) | MPR | Notes |
|---|---|---|---|---|---|---|---|---|
| AXIAL | DWI | EPI | 3 | 0.3 | YES | 23 cm | no | Angle to Corpus. Cover Skull Base to Vertex. Send only B1000 & ADC. |
| SAG | T1 | 3D TFE | 1.2 (160-170 slices) | 0 | no | 24-25.6 cm | no | Send to NQ_Epilepsy. See specific NeuroQuant 3D T1 scan parameters above. |
| COR OBLIQ | T2 STIR | 2D STIR TSE | 2 | 0.2 | YES | 23 cm | no | Whole brain, perpendicular to temporal lobe. |
| COR OBLIQ | T2 | 2D TSE | 2.5 | 0.25 | no | 15 cm | no | Small FOV centered on hippocampi. Cover all of hippocampi, from optic chiasm to splenium of corpus callosum. |
| AXIAL | SWI | 3D GRE | 2 | -1 | no | 23 cm | no | Angle to Corpus. Cover skull base to vertex. Ok to add slices. |
| Contrast injection | ||||||||
| SAG | T2 FLAIR | 3D IR-TSE | 1 mm | 0 | YES | 23 | AXIAL, COR OBLIQ | Angle to interhemispheric fissure. Cover ears and nose. Spacing and gap are variable. |
| SAG | T1 | 3D TSE | 1 mm | 0 | YES | 23 | AXIAL, COR OBLIQ | Angle to interhemispheric fissure. Cover at least 1cm above vertex through skull base. Cover entire nose. |