About the Portland Alcohol Research Center: The PARC
Lay Summary of Center Activities

The National Institute on Alcohol Abuse and Alcoholism (NIAAA) of the National Institutes of Health (NIH) has established the Portland Alcohol Research Center (PARC) at the VA Portland Health Care System (VAPHCS) and the Oregon Health & Science University (OHSU). The PARC has been continuously funded by NIAAA since 1996 to support alcohol-related research and is currently funded through 2025.
During the current funding period, the center has supported the coordinated research efforts of 14 scientists at the VA and OHSU, in areas ranging from behavioral neuroscience to molecular biology. More than 30 people are working on different aspects of the research. The PARC is one of 22 alcohol research centers funded by NIAAA, which is the primary source of all funding for alcohol research in the United States. Dr. Tamara Phillips is the center Director.
Overview: Uncovering the Genetics of How the Brain Adapts to Alcohol
Two overarching themes have historically united the work of the Portland Alcohol Research Center (PARC). The first is genetic risk factors that contribute to the etiology of alcohol use disorders. The second is the role of genetic factors in the consequences of alcohol use. Cumulative, coordinated efforts of the projects and cores generated the current overall center topic under investigation, which is The Role of the Tetrapartite Synapse in Risk for and Response to Alcohol Drinking. The tetrapartite synapse is composed of the pre- and post-synaptic membranes, astroglial processes and interstitial extracellular matrix. Alterations in any of these components are expected to alter synaptic function and thus, have the potential to impact risk for alcohol use and responses to alcohol. A major strength of the PARC is the inclusion of both rodents and non-human primates in the research, which results in more comprehensive analysis and provides important translational links.
A significant focus of our research is on individual differences in voluntary alcohol intake and brain region-specific gene expression differences in mice and monkeys corresponding with intake differences. In our current work, we focus on genetic influences on level of chronic alcohol use and neuroadaptation in mice and monkeys. The genetic risk and protective markers, patterns of neural circuitry differences and changes, and gene network information will ultimately help us and others to develop strategies for the prevention and treatment of alcoholism. Our existing genetic and imaging network analyses in mice and/or non-human primates have provided important information about relevant brain regions and pathways, and led to our focus on the tetrapartite synapse.
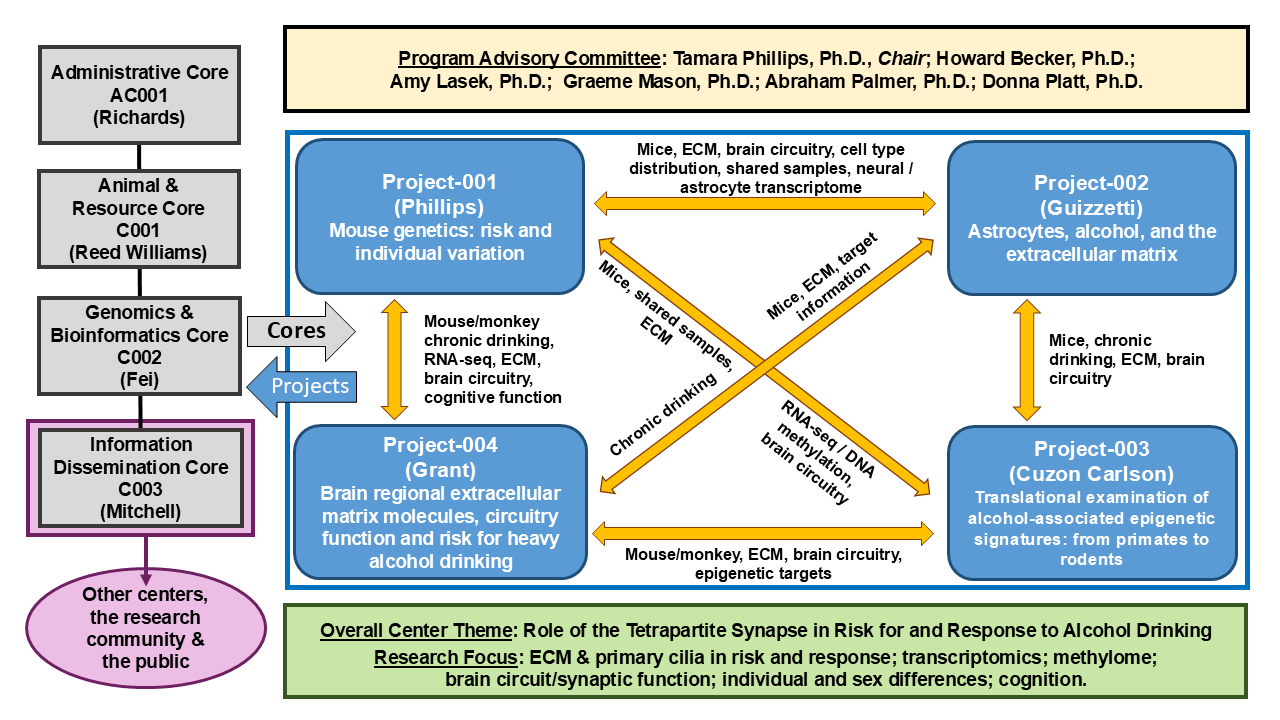
Four research Projects (P001-P004) and two service Cores (C001 and C002) address the PARC themes, using mouse models and non-human primate models. An outreach Core (Information Dissemination Core, CD001) continues to train pre-doctoral, post-doctoral and medical students in alcohol research, disseminate research findings to the public, and engages in a range of educational activities with elementary to high school students. An Administrative Core (AC001) provides scientific, organizational and budgetary oversight, makes executive decisions as required, and assists with coordination of Information Dissemination Core educational and science sharing activities. The components are summarized in the figure below.
Our bioinformatics efforts have enabled expansion of a key strength of our group from the analysis of the contributions of individual genes on behavioral functions of the whole organism to include gene network identification and extend our analyses to considerations of both coexpression and cosplicing, as well as differential wiring and differential variability; this continues in the Genomics and Bioinformatics Core (C002). Our Animal & Resource Core (C001) generates mice and collects behavioral data to provide consistency for all components of the center. In addition, the PARC has an extensive network of collaborations with alcohol researchers at other institutions, including those linked to other Centers and consortia. The figure shows the current PARC components, Leads, Program Advisory Committee members and interactions. In summary, the PARC seeks to identify genetic factors that place individuals at risk or protect them from alcoholism, and to identify genetic and neural consequences of excessive alcohol use, with a particular focus on those genetic factors that contribute to changes in synaptic function relevant to alcohol addiction. This knowledge will facilitate intervention and development of better therapeutics to treat alcohol use disorders.