Research
Cardiovascular Computational Mechanics Research
Mechanical forces play a significant role in biology. For example, blood flow forces regulate cardiovascular remodeling and heart development and affect thrombus growth, and surface tension forces determine the spread and transport of pulmonary surfactant inside the lungs.
Research projects at the Rugonyi Lab aim to understand the role of biomechanical forces on heart development, mural thrombogenesis and pulmonary surfactant function. To this end we use a combination of in vivo physiological data, imaging, mathematical modeling and computational simulation. Our ultimate goal is to understand the biological mechanisms by which mechanical forces affect biological processes.
Read more about our research projects by expaning the sections below.
Rugonyi Lab research news
The Rugonyi Lab's work on cardiac development was featured in the July 2013 issue of International Innovation, an international publication dissemination resource. International Innovation, accessible through Research Media, is a not just a publication for scientists, but the wider scientific, technology and research communities, dedicated to promoting discoveries and innovations in research and technology globally.
Projects
-
Cardiac growth and early developmental stages
The objective of this project is to better understand the origins of congenital heart disease (CHD), which affects about 1% of newborn babies in the U.S. and is the leading non-infectious cause of death among infants. The Rugonyi Lab is interested in the role of hemodynamic forces (forces exerted on tissues by the flow of blood) on cardiac development. In animal models, perturbations of the normal flow of blood through the heart during embryonic development lead to cardiac defects that resemble those found in humans. While genetic defects are known to underlie some cardiac malformations, abnormal hemodynamic conditions are just as likely to be responsible for many heart defects observed in human babies.
Although perturbed hemodynamic forces are known to lead to CHD, the mechanisms by which this happens are not fully understood. This is in part due to the complexity of the interactions between cardiac tissue, blood flow and cellular responses to mechanical stimuli; and in part due to the many technological challenges associates with measuring forces and deformations on small hearts that are beating fast.
Our goal is to use a combination of engineering and biology tools to unravel the mechanisms by which hemodynamic forces affect heart formation. To this end, we alter blood flow conditions during early embryonic cardiac development. We use chicken embryos because they are easy to access and manipulate, while genetic processes are highly conserved among vertebrate species. We use state-of-the-art imaging techniques to visualize the response of tissues to altered blood flow conditions.
Our experiments are complemented by computational investigations of blood flow and wall stresses in the developing heart, both under normal and abnormal hemodynamic conditions. The combination of both experimental and computational approaches enables us to shed more light towards understanding development of the heart and CHD.
We have obtained valuable data on blood flow velocities, blood pressures, histological and ultrastructural composition of the developing heart. Together, our data suggests that blood flow is a strong determinant of cardiac tissue growth and remodeling, as well as development of congenital heart defects.
Sample results
4D reconstruction of cardiac outflow tract
Beating heart outflow tract of a day 3 chick embryo (HH18). Segmented from 4D optical coherence tomography (OCT) images.
Blood flow through normal and banded embryos
The above video shows blood flowing through the outflow tract (OFT) of a normal embryo at day 3 (HH18). The OFT is roughly 800 microns in length.
The above video shows blood flowing through an embryo after undergoing outflow tract banding (OTB) at day 3 (HH18). The band can be seen across the middle of the outflow tract in black. Banding increases resistance to the normal flow of blood, thereby altering normal cardiac development. This provides valuable insights into some of the possible causes for CHD.
Comparing the endocardium of normal and outflow tract banded embryos
Normal HH24 chick embryo endocardium, displaying each cell as a different color. Focused ion beam scanning electron microscopy images were acquired through the endocardium layer of the outflow tract, and cellular tissue was segmented to reconstruct 3D cellular structures.
HH24 chick embryo endocardium after surgical manipulation, displaying each cell as a different color. Outflow tract banding was used to alter hemodynamics for 24 hours before tissue collection. Focused ion beam scanning electron microscopy images were acquired through the endocardium layer of the outflow tract, and cellular tissue was segmented to reconstruct 3D cellular structures.
Blood flow dynamics within the cardiac outflow tract (OFT)
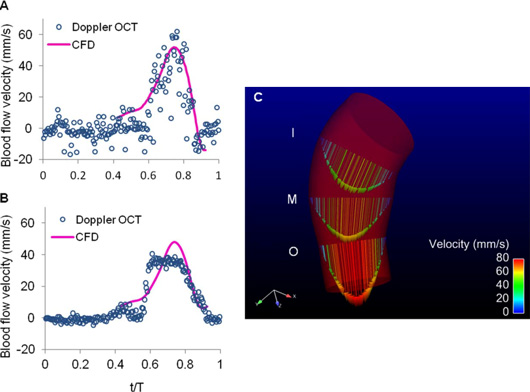
Blood flow dynamics within the cardiac outflow tract (OFT) of the representative 3-day (HH18) chick embryo. (A) and (B) Comparison of centerline velocities measured with Doppler optical coherence tomography (OCT) and calculated with our Computational Fluid Dynamics (CFD) model (C) CFD model of the heart OFT at maximal wall expansion, showing velocity profiles at three cross-sectional planes: I, M and O.
Published as: Figure 9 in Liu A, Yin X, Shi L, Li P, Thornburg KL, et al. (2012) Biomechanics of the Chick Embryonic Heart Outflow Tract at HH18 Using 4D Optical Coherence Tomography Imaging and Computational Modeling. PLoS ONE 7(7): e40869. doi:10.1371/journal.pone.0040869
-
An abdominal aortic aneurysm (AAA) is a pathological dilation of the abdominal aorta, which carries a high mortality rate if ruptured. The most commonly used surrogate marker of rupture risk is the maximal transverse diameter of the aneurysm. But maximal transverse diameter is not an accurate predictor of rupture risk.
The Rugonyi Lab is currently developing computational approaches to enhance patient-specific risk assessment and to follow AAA progression over time.
-
Cardiac dynamics segmented from 4D OCT images of the chick heart outflow tract at day 3 of incubation (cardiac cycle 370ms).
he goal of the Biological Shape Spaces project is to enable scientists to efficiently find relationships between the shape and function of biological systems, and to better understand how alterations in shape can alter function and vice-versa.
For biological systems, shape (i.e., the morphology of an organ or organism) is frequently retrieved in the form of images, which then need to be processed and interpreted. Advances in imaging technologies are allowing an ever-increasing amount of image information with increasing resolution. Despite these advances, the analysis of images in Medicine and Biology remains largely manual, and thus time consuming and subject to operator interpretation. Quantitative, functional interpretation of shape variations remains a fundamental challenge. Computational algorithms to analyze images and shape, while available, are often ad hoc, not easily generalizable and thus usually difficult to optimize for specific applications, and frequently out of the reach of non-experts.
Advances in high-throughput imaging have led to the rapid accumulation of shape information, but the tools to analyze these data have not kept pace. The lack of a coherent framework for quantifying and analyzing biological shape has prevented the objective testing of many hypotheses that rely on morphological data. Our inability to systematically link shape to genetics, development, function, environment, and evolution has frustrated advances in biological research across multiple spatial and temporal scales, from understanding how environmental influences alter developmental morphology to interpreting adaptive responses and radiations in the paleontological record.
To breach this important gap, investigators from eight institutions, including OHSU, are collaborating in a project funded by NSF to develop the next-generation computational tools that will allow biologists and physicians to analyze and retrieve shape information automatically, and enable discovering of relationships between shape and function. The goal is to not only develop better tools for mathematically describing shape, but also methods for extracting biologically meaningful information on morphological variation. The group consists of experts in Biology, Engineering, Computer Science and Mathematics determined to develop and bring tools to the scientific community that will enable optimal and fast analysis of shape and large imaging files. The collaborating PIs in this project are: Dr. Washington Mio (lead PI) from Florida State University; Dr. Surangi Punyasena, from University of Illinois at Urbana-Champaign; Dr. Ge Yang, from Carnegie Mellon University; Dr. Rolf Mueller, from Virginia Tech; Dr. Cindy Grimm, from Oregon State University; Dr. Charless Fowlkes, from University of California at Irvine; and Dr. Sandra Rugonyi, from the OHSU Department of Biomedical Engineering.
The collaborative project was launched in November 2010, with the First Workshop on Quantification of Biological Shape, organized by Dr. Rugonyi, and hosted at OHSU. The group has developed a website (see bioshapes.org) that will facilitate search and retrieval of images and image shape data, as well as sharing developed computational tools to find meaningful relationships between biological shape and function.
-
Focused ion beam scanning electron microscopy (FIB-SEM)
Focused ion beam scanning electron microscopy (FIB-SEM), or "slice and view," allows 3D imaging of en bloc biological samples with nanometer resolution. During image acquisition, the FIB mills thin sections of about 4 nm thick from the block, and then the SEM images the block face. This is done repeatedly to obtain a 3D image of the sample. In our work we use the FEI Helios 650 NanoLab™ DualBeam™ system from the OHSU Multiscale Microscopy Core. We image cardiac tissues from chicken embryos during early heart development.
Endocardium
Slice-and-view image of the HH24 chick embryo heart outflow tract endocardium. The endocardium is a monolayer of endocardial cells that is in contact with intracardiac blood flow.
Cardiac jelly
Slice-and-view image of the HH24 chick embryo heart outflow tract cardiac jelly layer. The cardiac jelly is predominantly an extracellular matrix layer in between the contractile myocardium layer and the endocardium layer.
Myocardium
Slice-and-view image of a portion of the HH24 chick embryo heart outflow tract myocardium. The myocardium is a compact layer of myocardial cells that actively contracts to pump blood through the circulation. The myocardium is also in contact with the cardiac jelly, and portions of the cardiac jelly can be observed in the video below.
-
Thrombosis and hemostasis
Thrombogenesis is a natural process by which the body acts on an injury to stop bleeding and heal the injured tissues. However, a growing thrombus may block a blood vessel preventing blood from reaching vital organs. Further, parts of a thrombus may detach and travel through the circulatory system causing heart attack and stroke. While the roles of cells and molecules that contribute to thrombus formation have been identified, and the steps that lead to thrombus formation, the so-called coagulation cascade, are fairly well known, the effect of blood flow on thrombus formation and transport of thrombogenic products is less understood.
Thrombus formation on thrombogenic surfaces has been shown to consist of 3 phases: (I) initial, slow growth; (II) linear growth; (III) plateau, during which the thrombus stops growing. Phase three may not be observed if the thrombus occludes the vessel. While phases one and two are somewhat understood, the mechanisms by which the thrombus stops growing are not well understood. Several factors, such as blood and thrombus-surface chemistry as well as blood flow might play a role. The Rugonyi Lab focuses on understanding the effect of blood flow on thrombus growth, with the ultimate goal of elucidating the mechanisms by which thrombi break lose or stop growing.
Pulmonary surfactant biophysics
Pulmonary surfactant is essential for normal breathing. The lack of sufficient amount of mature surfactant causes respiratory distress syndrome (RDS) in premature infants, and can worsen the patient condition in adults with acute respiratory distress syndrome (ARDS). In the lungs, pulmonary surfactant forms a thin surface film, generally believed to be a monolayer, at the interface between air and a thin liquid layer that coats the alveoli. This surfactant film reduces surface tension, a force that tends to collapse the alveoli causing lung injury.
In situ experiments demonstrate that surface tension reaches very low values (~1 mN/m) during exhalation. However, these very low values are difficult to reproduce in vitro. Unlike in the lungs, during in vitro compression of pulmonary surfactant monolayers, surface tension decreases until the monolayer starts to thicken forming multi-layer films, process that is usually referred to as monolayer collapse and that occurs at a constant surface tension (the collapse or equilibrium surface tension, ~ 24 mN/m). Why pulmonary surfactant film responds differently to compression in situ and in vitro is not well understood. Dr. Rugonyi's research has focused on the kinetics of surfactant collapse - the formation of multi-layers from a monolayer film - and how collapse rates are affected by (I) the rate of film compression; (II) the film surface tension; and (III) the sub-phase liquid thickness. The ultimate goal is to understand the mechanisms by which proper surfactant function is achieved.