Projects
Protein ubiquitination is arguably the most complex and widespread signaling network in biology. Unlike post-translational modifiers such as phosphorylation or acetylation that are binary, ubiquitination can be highly customized through further post-translational modification of ubiquitin itself. It is this unique feature that allows ubiquitin and related ubiquitin-like modifiers to regulate diverse aspects of biology including proteasomal degradation, DNA damage response, and immunity in a highly controlled and specific fashion.
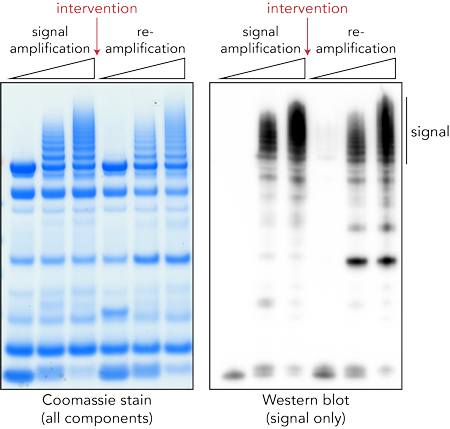
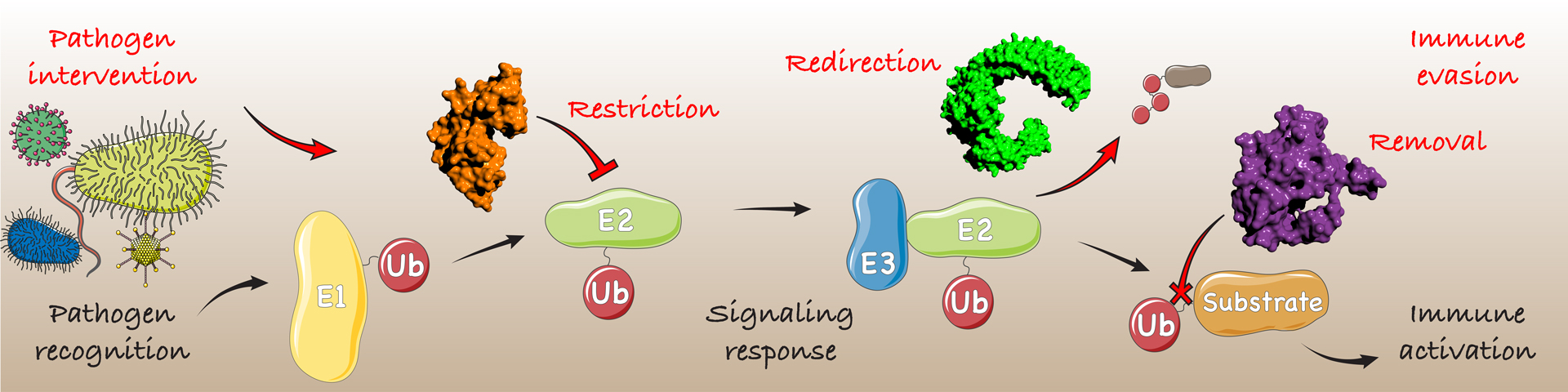
Ubiquitination is tightly controlled by hundreds of regulatory enzymes (e.g. E1, E2, E3) and breakdown of this system is implicated in numerous human diseases. In addition, its roles in regulating immune signaling place ubiquitin in the crosshairs of microbial intervention during infection. Pathogens such as bacteria or viruses encode effector enzymes with the specific purpose of interfering with host ubiquitin signaling. These enzymes can restrict, redirect or remove ubiquitin signals, in some cases through chemistries that are entirely foreign to the human system.
Our lab seeks to identify examples of microbial intervention in ubiquitin signaling, understand their modes of action, and assess their contributions to infection and disease. Our ability to biochemically reconstitute ubiquitin signaling in vitro allows us to witness the intervention firsthand in a controlled setting. We use biophysical and structural techniques such as X-ray crystallography to visualize these host-microbe interactions on the molecular level. Coupled with cellular models of infection, we identify the biochemical mechanism and cellular context of the intervention. By this approach we anticipate our work will unveil many exciting new aspects of infection and immunity.