Research
Nan Laboratory Research Overview
The main theme of research in the Nan lab is nanoscale spatial systems biology. Our goal is to understand how biological molecules and their functional assemblies work in their physiological contexts across the scales ranging from single molecules (nanometers) to multicellular structures (millimeters). In particular, we are interested in how oncogenic signaling of the Ras small GTPases and HER family receptors are regulated in space and time in individual cells, and how variations in the molecular interactions in individual cells are linked to heterogeneities in phenotypes and fates in cell populations. To these ends, we invest major efforts in developing single-molecule and superresolution imaging techniques, and we take a multidisciplinary approach that combines imaging, computation, and molecular and cell biology in our research.
-
Fluorescence microscopy has been instrumental to biomedicine but is intrinsically limited in spatial resolution (to ~200 nm) by the diffraction of light. In the last two decades or so, a growing list of imaging techniques have been introduced to circumvent the diffraction limited resolution – hence the term ‘superresolution microscopy’ (SRM) [1,2]. A subset of existing SRM techniques is based on sub-diffractive localization of single molecule emitters and commonly referred to as ‘Single-molecule localization microscopy’ [3] (SMLM, Figure 1). The best-known examples of SMLM include photoactivated localization microscopy (PALM) [4,5], stochastic optical reconstruction microscopy (STORM) [6], point accumulation in nanoscale topography (PAINT) [7] and its recent derivation DNA-PAINT [8].
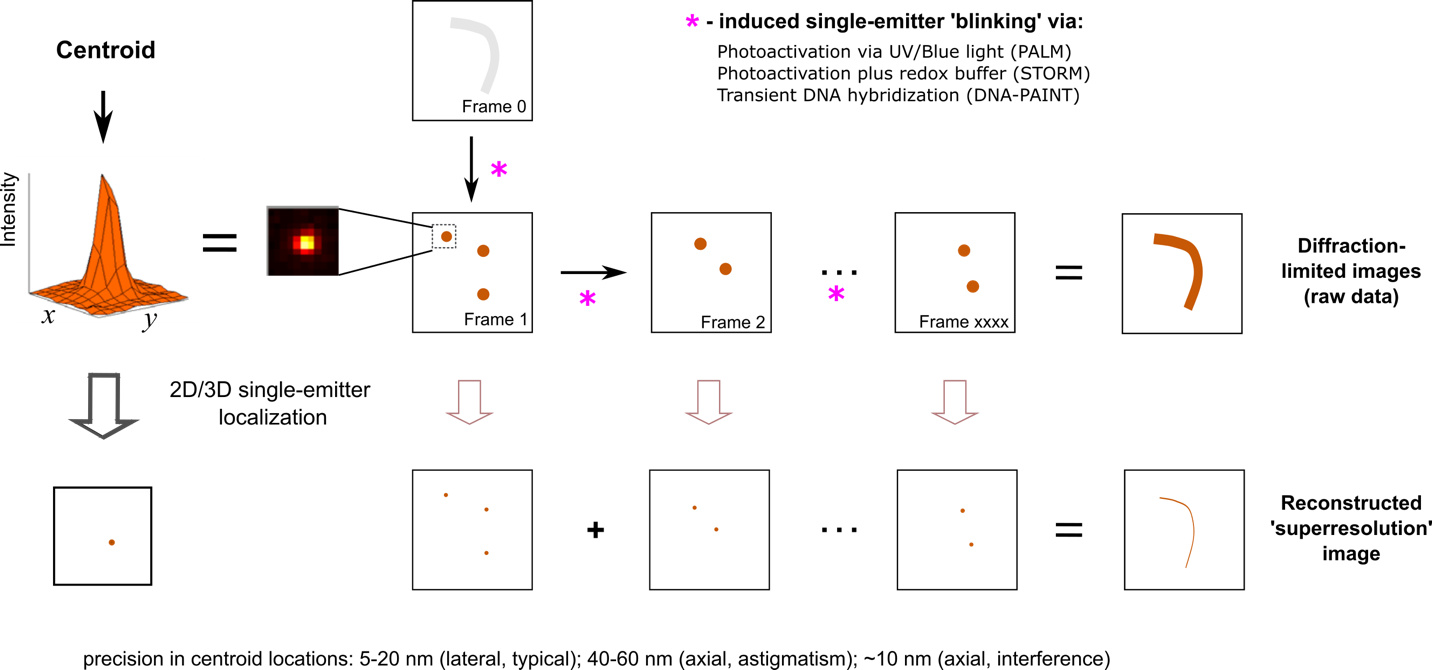
Figure 1. Superresolution microscopy (SRM) based on single-molecule localization. All SMLM-based SRM techniques – PALM, STORM, and (DNA-)PAINT – employ mechanisms for efficient single-molecule ‘blinking’, such that signals from single emitters are spatially separated for precise localization. Otherwise, the densely packed fluorophores (as in most biological staining) would emit fluorescence signals all at once to yield a diffraction-limited (‘smeared’) image. PALM and STORM take advantage of photoswitching of certain fluorescent probes, such as photoactivatable fluorescent proteins (e.g. PAmCherry and mEos) [9] or small molecule organic dyes (e.g. AlexaFluor 647) [10] in combination with specialized imaging buffers. In PAINT, single-molecule localization events arise from transient binding events that lead to detectable fluorescence, for example when Nile Red molecules bind to lipid bilayers [7]. Nile red is barely fluorescent when dissolved in water but turns fluorescent when inserted into a hydrophobic lipid bilayer [11]. In DNA-PAINT, transient hybridizations between a pair of short, complementary oligonucleotides yield bursts of detected fluorescence events [8]. In all cases, a ‘super-resolved’ image can be reconstructed by combining the locations of all the detected single molecule events.
For each fluorescent molecule, it’s lateral (x, y) position can be localized to a precision roughly equivalent to the half width of the diffraction limited image (typically 120-160 nm depending on the wavelength and lens aperture) divided by the square root of the number of detected photons [12]. Considering mechanical drift, pixelation, and other factors, each fluorescent molecule could be localized to ~10 nm or better precision with as few as 500-1,000 photons collected. This translates into 20-25 nm spatial resolutions if the sample labeling is sufficiently dense (to achieve Nyquist sampling). In fact, common structures such as the microtubule network in cultured mammalian cells can be imaged at similar spatial resolutions using PALM, STORM, or DNA-PAINT (Figure 2). We use all three techniques in our research depending on the application.
![Figure 2. Superresolution imaging of microtubules in cultured cells with PALM (left), STORM (middle), and DNA-PAINT (right). The STORM image is taken from ref [13], and the DNA-PAINT image from ref [8].](/sites/default/files/2022-04/Superresolution%20Microscopy%20Research%20Fig%202.png)
Figure 2. Superresolution imaging of microtubules in cultured cells with PALM (left), STORM (middle), and DNA-PAINT (right). The STORM image is taken from ref [13], and the DNA-PAINT image from ref [8]. The image of a single fluorophore not just reports its lateral (x, y) position but also encodes its axial (z) information. Depending on its z position, the resulting image may be slightly different, but the difference can be small, making it difficult to accurately extract the z position from a ‘normal’ single-molecule image. One strategy for precise three-dimensional (3D) localization of single molecules is to introduce astigmatism, such that the single-molecules take very different shapes depending on their axial locations (e.g. below or above the focal plane). This can be achieved by inserting a cylindrical lens into the detection path (Figure 3, left), a relatively straightforward setup [14]. The astigmatism based strategy affords z-location precisions around 3x worse than in the x-y position (that is, 20-30 nm). More precise z-localizations (~10 nm) were achieved using single-photon interferometry (termed interferometric PALM or iPALM) [15] (Figure 3, right).
![Figure 3. Three-dimensional (3D) SRM based on optical astigmatism (left) or interferometry (right). Schematics and plots on the left were taken from ref [14], and those on the right taken from from ref [15].](/sites/default/files/2022-04/Superresolution%20Microscopy%20Research%20Fig%203.png)
Figure 3. Three-dimensional (3D) SRM based on optical astigmatism (left) or interferometry (right). Schematics and plots on the left were taken from ref [14], and those on the right taken from from ref [15]. Both 3D localization mechanisms are applicable to all SMLM-based SRM. For example, we have used STORM with both astigmatism based 3D localization (Figure 4, left) and with iPALM (Figure 4, middle). The iPALM has also been combined with DNA-PAINT for ultrahigh resolution 3D imaging of nanostructures such as caveolae in cells (Figure 4, right). The Nan lab hosts both setups for 3D SRM (Thanks to Drs. Rainer Daum and Meike Pederson for helping acquire a commercial iPALM prototype).
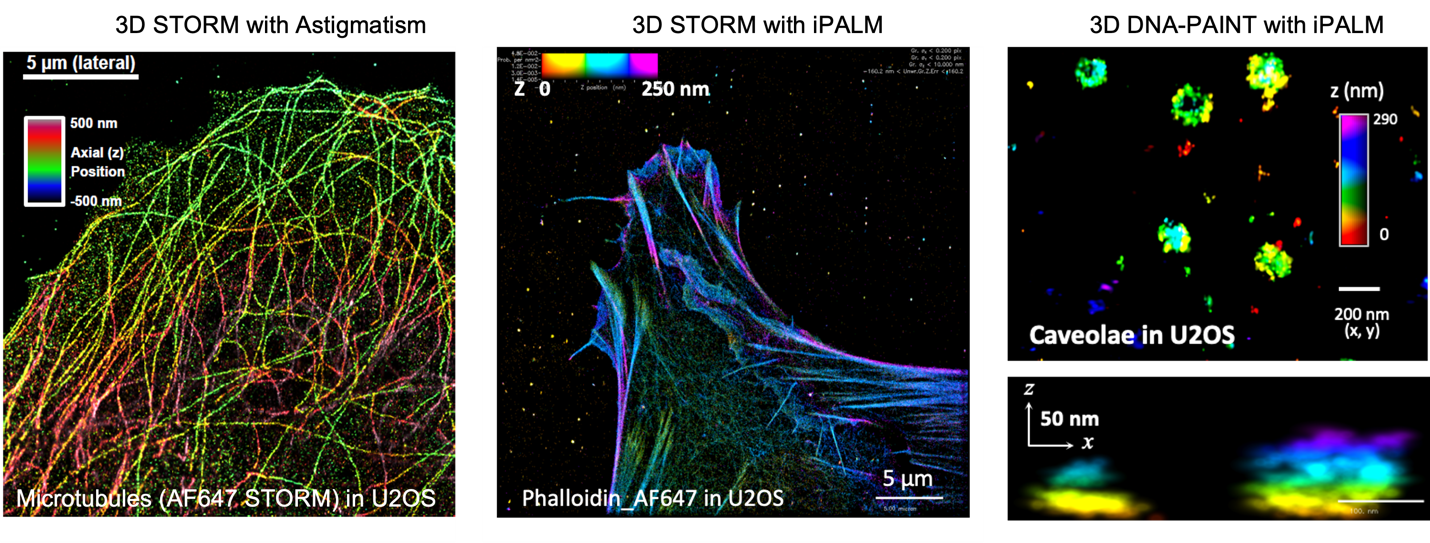
Figure 4. 3D-SRM imaging of cellular structures. Left, 3D-STORM image of microtubules in U2OS cells based on astigmatism. Middle, 3D-STORM image of actin filaments in U2OS cells taken with iPALM. Right, 3D DNA-PAINT image of caveolae in U2OS cells taken on an iPALM. References
- Huang, B. Super-resolution optical microscopy: multiple choices. Curr. Opin. Chem. Biol. 2010, 14, 10–14.
- Sigal, Y.M.; Zhou, R.; Zhuang, X. Visualizing and discovering cellular structures with super-resolution microscopy. Science 2018, 361, 880–887, https://doi.org/10.1126/science.aau1044.
- Patterson, G.; Davidson, M.; Manley, S.; Lippincott-Schwartz, J. Superresolution imaging using single-molecule localization. Annu. Rev. Phys. Chem. 2010, 61, 345–367, https://doi.org/10.1146/annurev.physchem.012809.103444.
- Betzig, E.; Patterson, G.H.; Sougrat, R.; Lindwasser, O.W.; Olenych, S.; Bonifacino, J.S.; Davidson, M.W.; Lippincott-Schwartz, J.; Hess, H.F. Imaging intracellular fluorescent proteins at nanometer resolution. Science 2006, 313, 1642–5, https://doi.org/10.1126/science.1127344.
- Hess, S.T.; Girirajan, T.P.K.; Mason, M.D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 2006, 91, 4258–72, https://doi.org/10.1529/biophysj.106.091116.
- Rust, M.J.; Bates, M.; Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 2006, 3, 793–5, doi:10.1038/nmeth929.
- Sharonov, A.; Hochstrasser, R.M. Wide-field subdiffraction imaging by accumulated binding of diffusing probes. Proc. Natl. Acad. Sci. U. S. A. 2006, 103, 18911–6, https://doi.org/10.1073/pnas.0609643104.
- Jungmann, R.; Avendaño, M.S.; Woehrstein, J.B.; Dai, M.; Shih, W.M.; Yin, P. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 2014, 11, 313–8, https://doi.org/10.1038/nmeth.2835.
- Shcherbakova, D.M.; Sengupta, P.; Lippincott-Schwartz, J.; Verkhusha, V. V Photocontrollable Fluorescent Proteins for Superresolution Imaging. Annu. Rev. Biophys 2014, 43, 303–29, https://doi.org/10.1146/annurev-biophys-051013-022836.
- Dempsey, G.T.; Vaughan, J.C.; Chen, K.H.; Bates, M.; Zhuang, X. Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging. Nat. Methods 2011, 8, 1027–1036, https://doi.org/10.1038/nmeth.1768.
- Greenspan, P.; Fowler, S.D. Spectrofluorometric studies of the lipid probe, nile red. J. Lipid Res. 1985, 26, 781–9.
- Thompson, R.E.; Larson, D.R.; Webb, W.W. Precise nanometer localization analysis for individual fluorescent probes. Biophys. J. 2002, 82, 2775–83, https://doi.org/10.1016/S0006-3495(02)75618-X.
- Bates, M.; Huang, B.; Dempsey, G.T.; Zhuang, X. Multicolor super-resolution imaging with photo-switchable fluorescent probes. Science 2007, 317, 1749–53, https://doi.org/10.1126/science.1146598.
- Huang W, B.W.; Q Bates X W, M.Z. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science (80-. ). 2008, 319, 810–813, https://doi.org/10.1126/science.1153529.
- Shtengel, G.; Galbraith, J. a; Galbraith, C.G.; Lippincott-Schwartz, J.; Gillette, J.M.; Manley, S.; Sougrat, R.; Waterman, C.M.; Kanchanawong, P.; Davidson, M.W.; et al. Interferometric fluorescent super-resolution microscopy resolves 3D cellular ultrastructure. Proc. Natl. Acad. Sci. U. S. A. 2009, 106, 3125–3130, https://doi.org/10.1073/pnas.0813131106.
-
(This section is being rewritten and will be updated soon)
-
(This section is under development; updates to come soon after our manuscript is made public)
-
(This section is being rewritten and will be updated soon)
-
(This section is being rewritten and will be updated soon)
-
(This section is being rewritten and will be updated soon)
Research Sponsors
Research in the Nan Lab is funded by these sponsors:
Knight Cancer Institute; Damon Runyon Cancer Research Foundation; FEI; National Cancer Institute, and the M.J. Murdock Charitable Trust.

Keywords: biological nanoscopy, single-molecule biophysics, cancer medicine