Research Interests
The major goals of our laboratory are to comprehensively understand the biology of recombinant adeno-associated virus (AAV) and to develop new AAV vector-mediated gene and cell therapies to treat various human diseases. Scientific curiosity is the most important, major driver of the lab, having led to many discoveries and breakthroughs in AAV vectors and gene therapy. We also fully enjoy leveraging our knowledge and expertise around AAV, virus and gene therapy research to understand broader and more fundamental biological questions that have yet to be addressed or answered in various scientific disciplines.

AAV & AAV Biology
AAV and AAV Biology
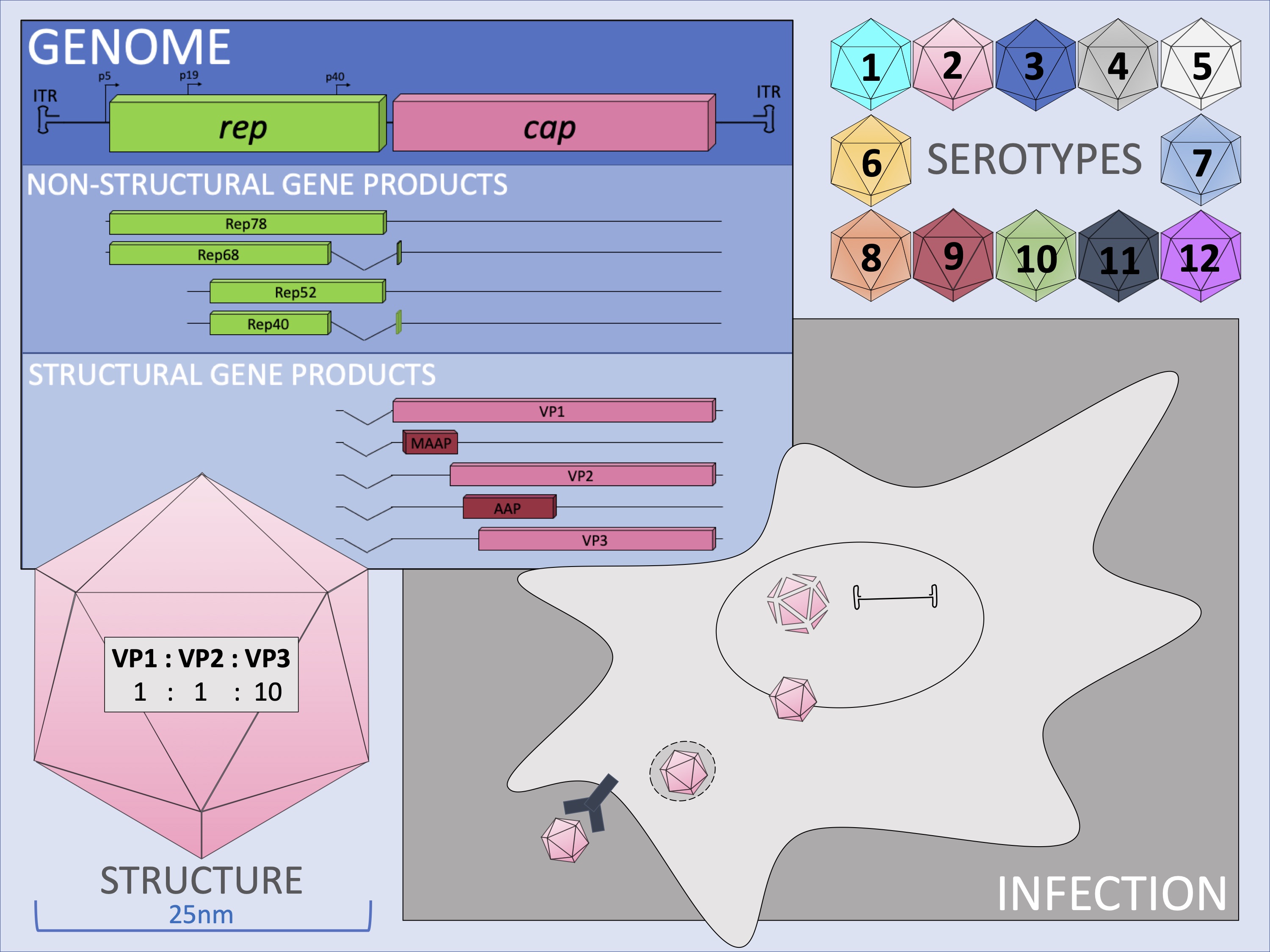
AAV belongs to the parvovirus family and is the smallest virus that has the simplest structure. The virus is 25 nm in diameter and is composed of a protein shell, called capsid, and a 4.7-kb single-stranded DNA genome coding two viral genes, the rep and cap genes. The rep gene encodes viral non-structural proteins, Rep40, Rep52, Rep68 and Rep78, translated from a single open reading frame (ORF), and play roles in viral genome replication and packaging. The cap gene encodes structural proteins VP1, VP2 and VP3 translated from a single ORF and two non-structural proteins, AAP (assembly activating protein) and MAAP (membrane-associated accessory protein) from +1 frameshifted ORFs. VP proteins form the viral capsid made of 60 VP subunits at a ratio of VP1(1):VP2(1):VP3(10). The AAV genome has inverted terminal repeats (ITRs) at both genome termini that form a secondary hairpin structure as a result of their palindromic sequences. AAV infects humans but does not cause any human diseases; therefore, AAV is nonpathogenic and offers an attractive vehicle to deliver therapeutic genes to human body.
AAV infects cells via cell surface viral receptors, enter cells, pass the nuclear envelop and disassemble in the nucleus, releasing the viral genome and establishing latent infection in non-dividing cells. In replicating cells, AAV viral genomes do not replicate or minimally replicate and therefore are lost eventually. A small portion of the viral genomes integrate into the host cellular DNA. In AAV serotype 2 (AAV2), viral genomes integrate into the AAVS1 site on human chromosome 19 in a site-specific fashion by a Rep-mediated mechanism. However, the current view is that this is not likely the mechanism of AAV2 latent infection in the human body, and AAV viral genomes remain as episomes (extrachromosomal viral genomes) for a long period of time to maintain latency. Upon coinfection or superinfection with adenovirus or other helperviruses such as herpes simplex virus and cytomegalovirus, the AAV enters the productive viral life cycle, replicating viral genomes and producing progeny.
At least 13 AAV serotypes have been identified and hundreds of naturally occurring AAV variants have been isolated in various organs from humans and non-human primates. Each serotype and naturally occurring variant exhibits unique biological properties and show serotype/variant-dependent tropism during the infection process.
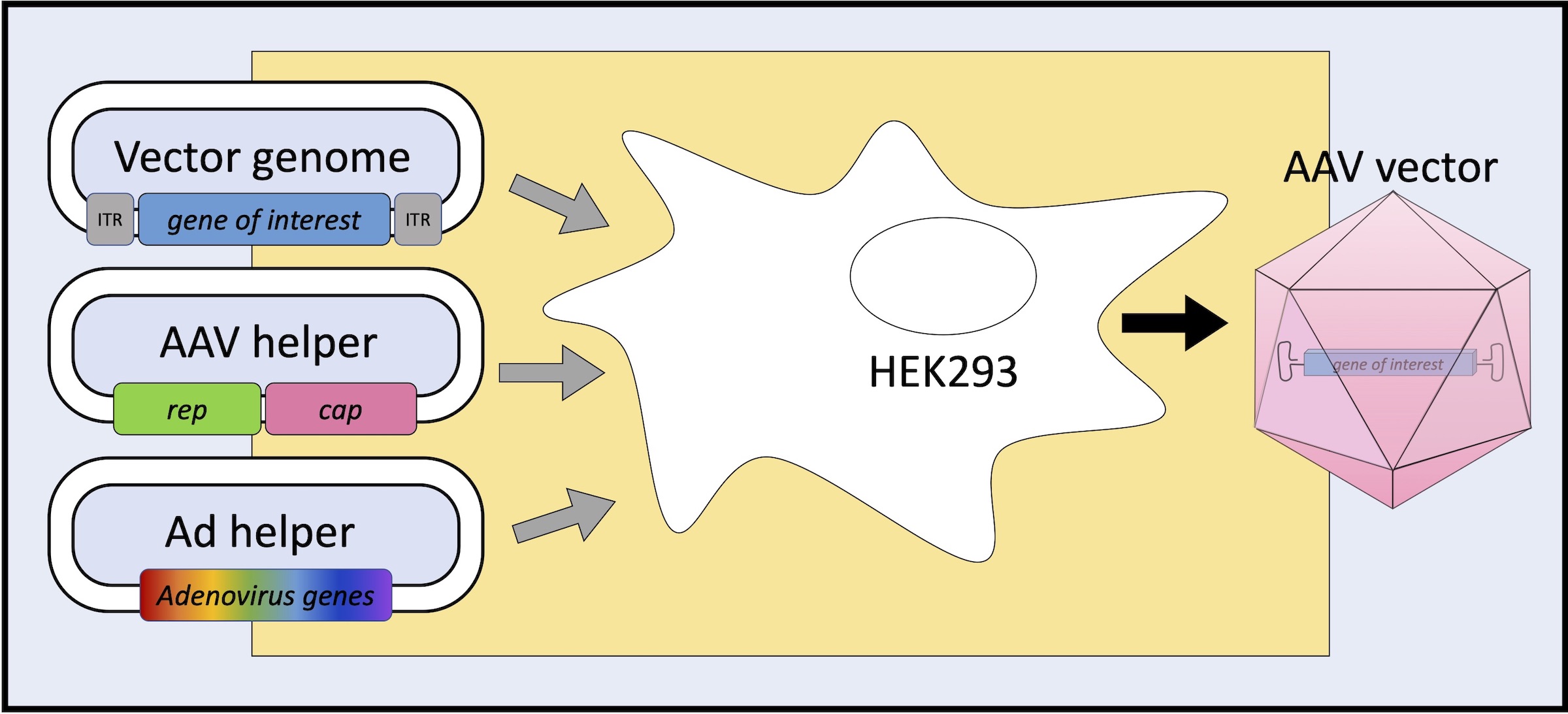
Cloning of the AAV2 genome into a plasmid backbone provided a model for understanding of the AAV virus life cycle. The discovery that the inverted terminal repeats (ITRs) were the only DNA sequences required in cis for AAV production in cultured cells led to the use of recombinant AAV (AAV) as a vector for gene therapy. In recombinant AAV vectors that deliver to cells a gene of interest (GOI) including therapeutic genes, the rep and cap genes are removed from the AAV viral DNA genome and replaced with GOI, leaving AAV ITRs as the only viral genome-derived component in recombinant AAV vector genome. AAV vectors can be produced in packaging cells (e.g., human embryonic kidney (HEK) 293 cells) by delivering AAV vector plasmid, AAV helper plasmid and adenovirus (Ad) helper plasmid to the cells by transient DNA transfection. The AAV vector plasmid provides an AAV vector genome to be packaged in the virion, the AAV helper plasmid supplies AAV Rep, VP and AAP proteins, and the Ad helper plasmid supplies adenovirus components essential for virus replication and production in cells.
Specific areas of interest
An understanding of basic AAV biology has been accomplished over five decades of studies, yet a number of complex questions remain unanswered, including how AAV capsids are formed, how AAV viral genomes are packaged into the capsid shell, how viral proteins and host proteins interact in the process of virion formation, how AAV enters the cells via receptor binding and endocytosis, traffics to the nucleus and uncoats releasing viral genome, how AAV viral genomes are processed via the host cell DNA damage responses and repair mechanisms, how viral capsid proteins, viral genome and host transcription/translation machinery interact enhancing and suppressing transgene expression, how capsid amino acid sequence and structure determine AAV biological properties, among others.
How did the overlapping AAP and VP ORFs co-evolve?
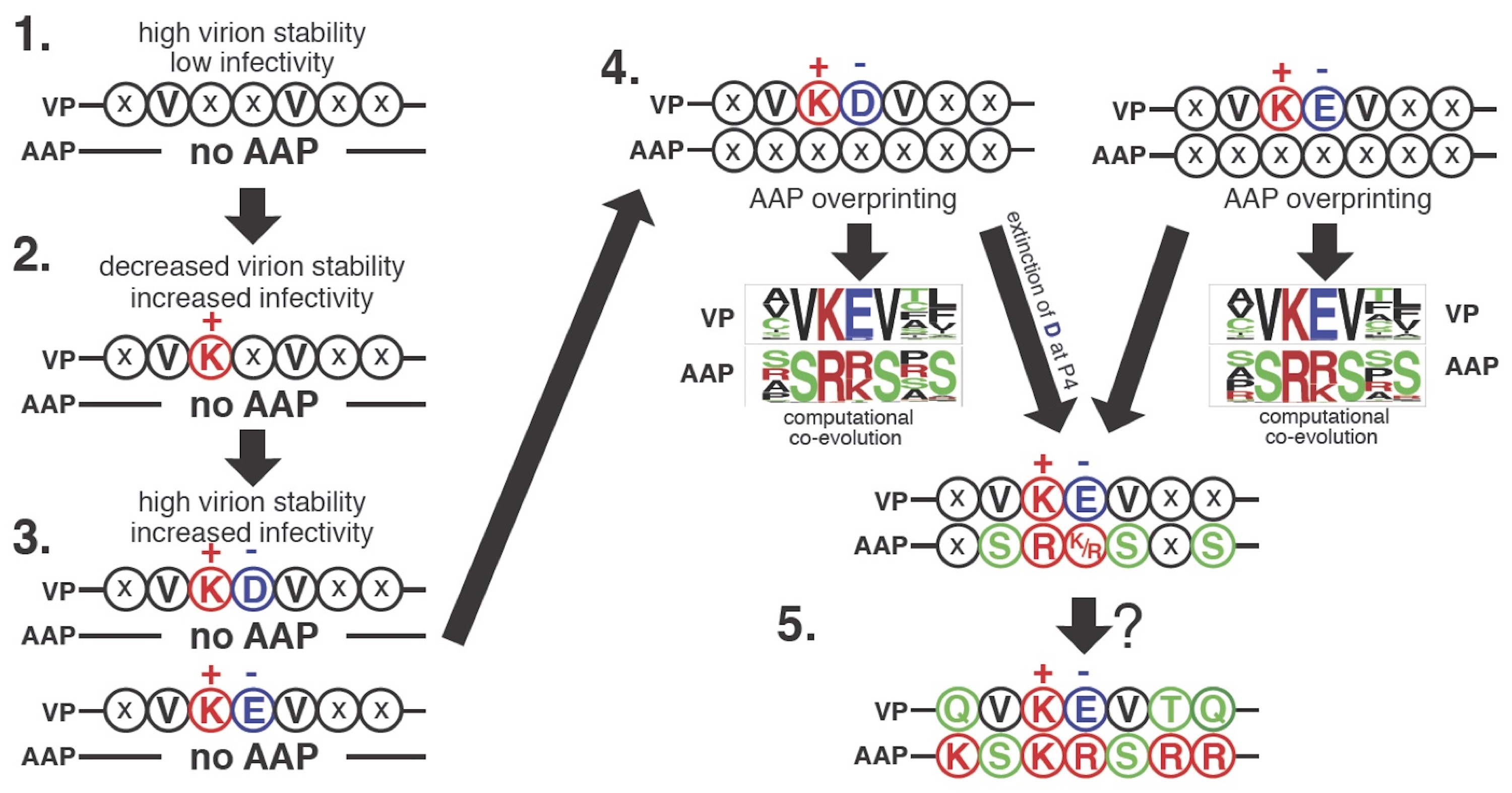
A proposed model of AAP2 ORF overprinting in the AAV2VP ORF deduced by evolutionary computation. The evolutionarily conserved QVKEVTQ and KSKRSRR motifs, a pair of overlapping heptapeptides in VP and AAP of AAV2, respectively, were randomized and experimentally evolved under no coevolutionary constraints, and a large number of capsid-forming VP mutants and functionally competent AAP mutants of these motifs were identified by the next-gen sequencing. The obtained heptapeptide information was then integrated into an evolutionary algorithm, with which VP and AAP were computationally co-evolved from random nucleotide sequences in silico. This experimental and computational evolution-based analysis revealed how overlap-evoked co-evolutionary constraints play a role in making the VP and AAP heptapeptide sequences into the present shape and led to a hypothetical model of VP and AAP co-evolution as depicted in this figure. The most primitive heptapeptide sequence in the AAV2 VP protein when no AAP existed is shown at the top left. The heptapeptide region in the present shape is shown at the right bottom.
Kawano, Y., Neeley, S., 1, Adachi, K., Nakai, H. (2013) An experimental and computational evolution-based method to study a mode of co-evolution of overlapping open reading frames in the AAV2 viral genome. PLoS One 8, e66211.
How does AAP contribute to capsid assembly?
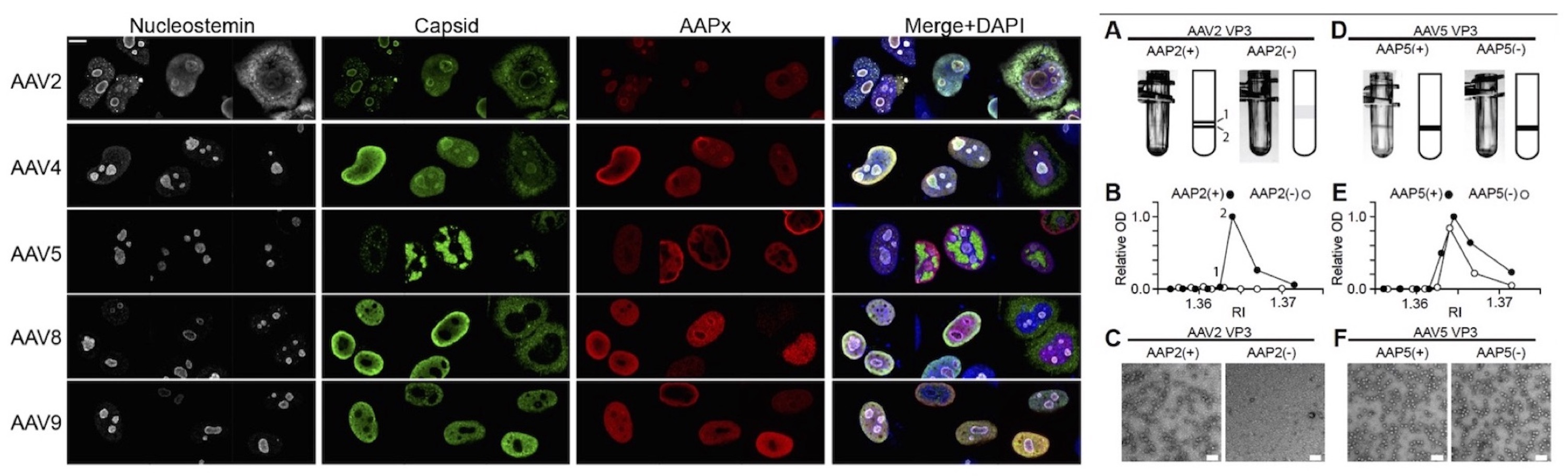
Heterogeneity of the roles of AAV in the process of AAV capsid assembly among different AAV serotypes. (Left Panel) Subcellular localization of AAP and assembled capsids in HEK293 cells. AAV2 and AAV4 assemble in the nucleolus while AAV5, AAV8 and AAV9 assemble in the nucleoplasm outside the nucleolus. (Right Panel) AAV VP3 only capsid production with or without AAP. AAV2 capsid formation requires AAP while AAV5 capsid formation does not.
Earley, L.F., Powers, J.M, Adachi, K., Baumgart, J., Meyer, N.L., Xie, Q., Chapman, M.S., Nakai, H. (2017) Adeno-associated virus assembly-activating protein is not an essential requirement for capsid assembly of AAV serotypes 4, 5 and 11. J. Virol. 91:e01980-16
How is the genome packaging capacity determined?
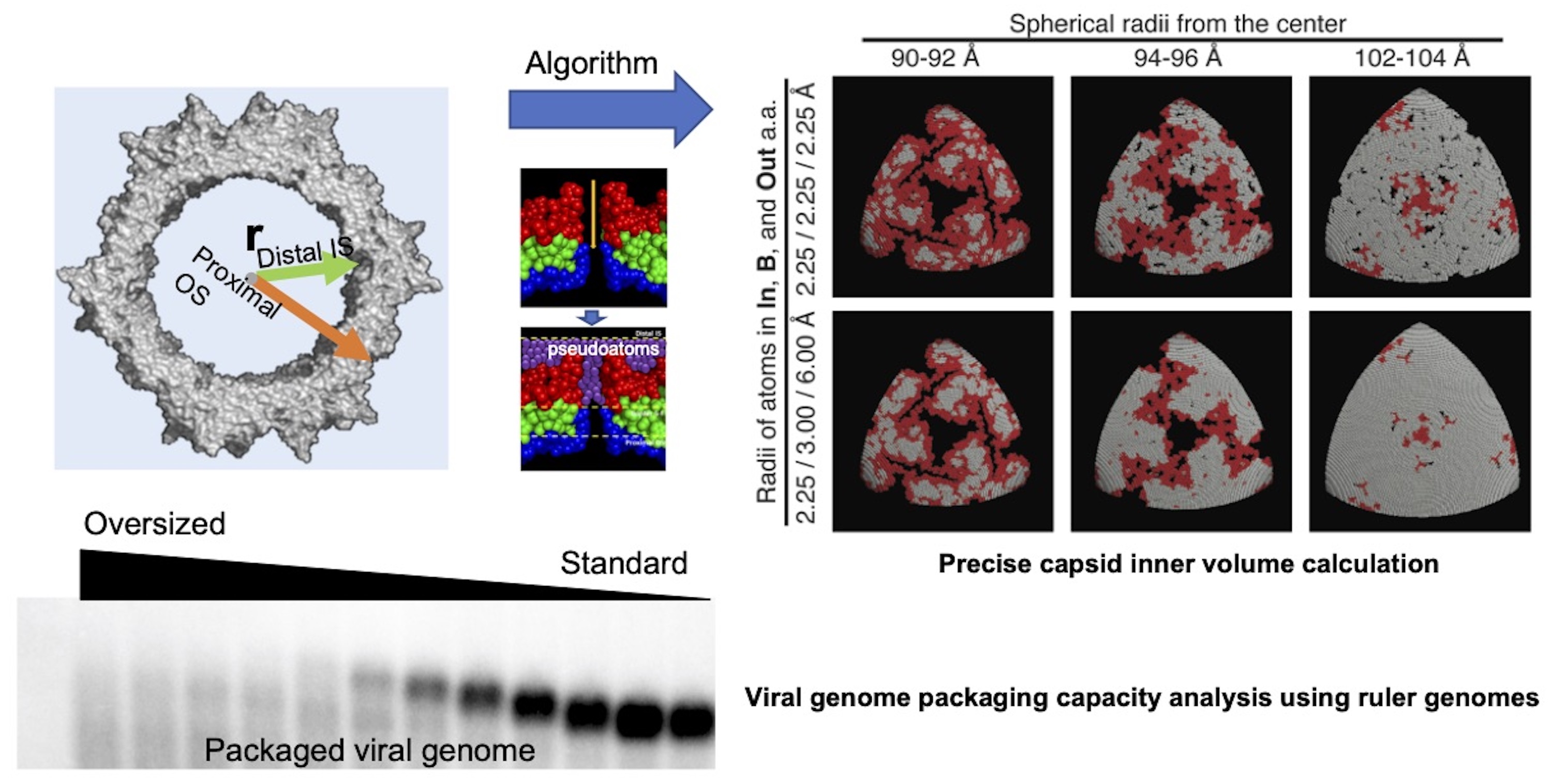
Methods of calculation of AAV capsid inner volume and determination of genome packaging capacity of the AAV capsid. (Top Panels) A computational algorithm has been developed to precisely calculate capsid inner volume. For this purpose, pseudoatoms and inflated atoms are used to flatten irregular outer surface of the capsid and fill interstices present inside protein molecules, respectively.
What cellular proteins interact with AAV proteins in the process of capsid assembly?
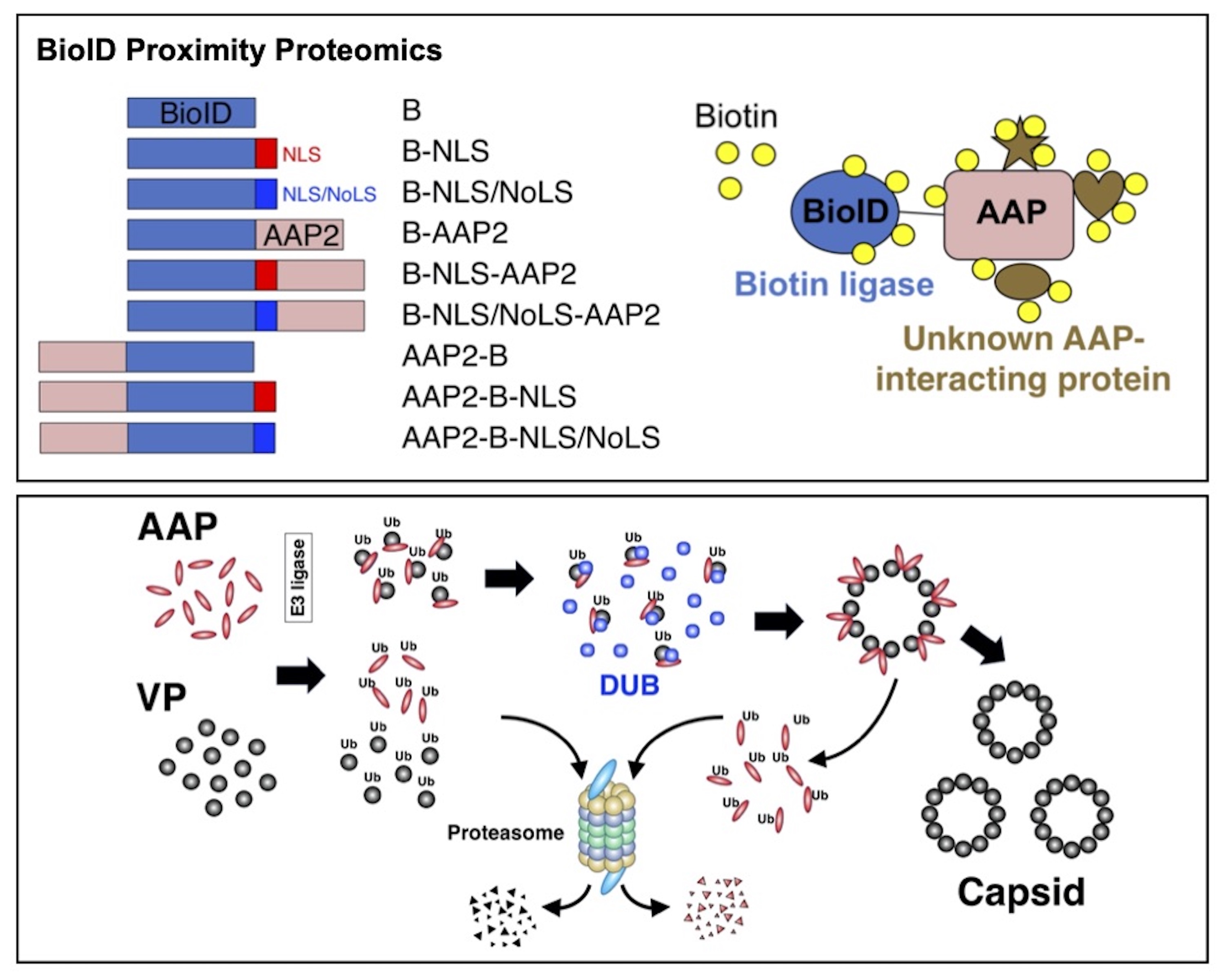
BioID approach to identify AAP-interacting cellular proteins (Top Panel). BioID proximity labeling followed by LC-MS/MS identifies AAP-interacting cellular proteins (e.g., deubiquitinating enzyme (DUB)). A preliminary hypothetical model of the interaction between DUB and AAV proteins in the process of capsid assembly (Bottom Panel).
How are the multifaceted biological phenotypes of AAV vectors governed in a system?
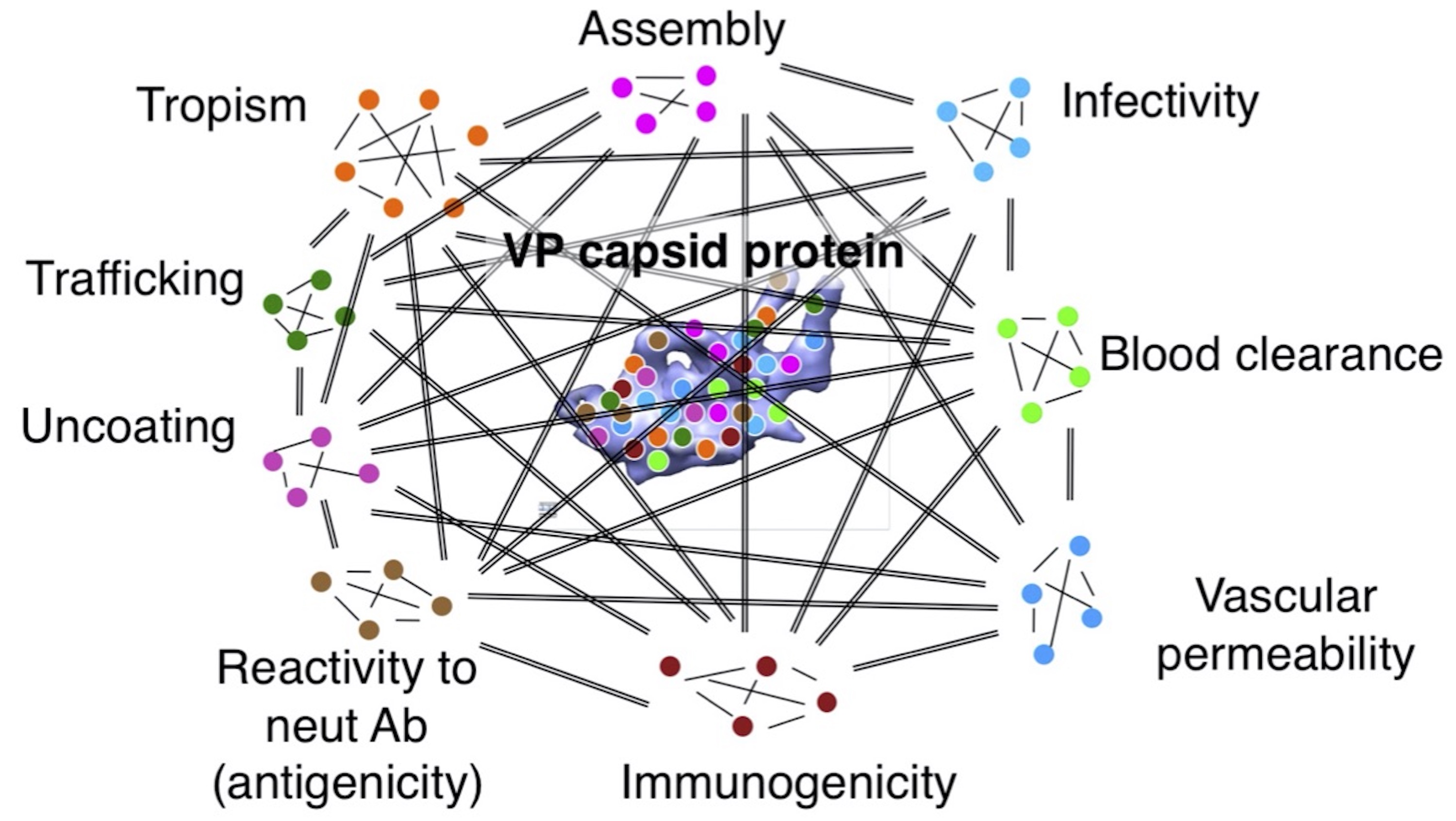
Multi-faceted biological functions of the AAV capsid protein. AAV capsid VP protein has multifaceted functions in the tertiary and quaternary structures and contains multiple functional modules and components that interact with each other in a system, leading to the manifestation of emergent properties. Note: this is how viruses can make the most of its limited structural components to maximize its functions. It is very challenging to understand the system that governs virus phenotypes by traditional approaches. We have been addressing this question by using data-driven systems biology approaches (see “Data-driven Research” tab).
Data-driven Research
Data-driven Research
Despite the simplest structure of AAV, the AAV vector biology and the underlying mechanisms of AAV vector-mediated gene delivery and transduction in cells are very difficult to understand. The AAV's biological phenotypes are primarily determined by the AAV capsid shell composed of VP proteins. To make a breakthrough in this field, the Nakai lab pioneered a virus barcoding approach to study AAV vectors and established AAV Barcode-Seq in 2010, a next-generation sequencing (e.g., Illumina sequencing)-based comprehensive phenotype characterization of hundreds of different AAV strains (AAV vectors derived from different serotypes, variants and mutants) in an unprecedentedly high-throughput manner. The AAV Barcode-Seq technology was first introduced to the gene and cell therapy community by Dr. Nakai in the Plenary Session at the Annual Meeting of the American Society of Gene and Cell Therapy 2011. Since then, the AAV Barcode-Seq has further evolved into more advanced technologies in various directions. In addition, the power of virus barcoding has become widely appreciated and the barcode-seq is currently employed by many laboratories. The Nakai lab continue to generate a massive amount of AAV capsid amino acid sequences - AAV biological phenotype relationship data, a small portion of which has been published in peer-reviewed journals and presented at scientific meeting. In addition to AAV Barcode-Seq, the Nakai lab generates large datasets pertinent to AAV vector biology by using other means and analyzes the data in various ways including machine learning. All the data will be eventually released to the public to maximize the utility of our dataset to advance research and gene therapy.
Specific areas of interest
Establishing novel data-driven approaches for AAV research
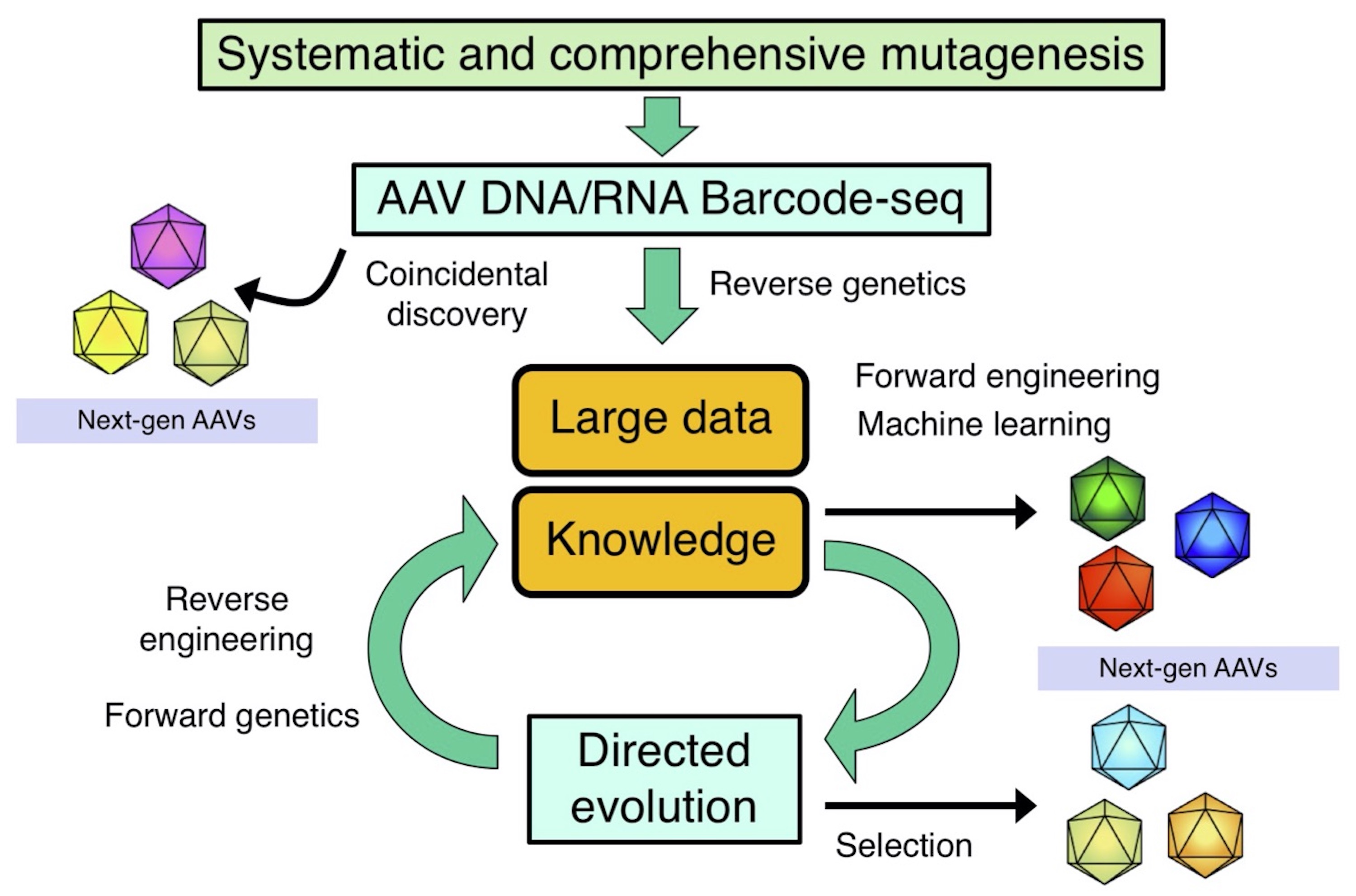
Our data-driven approach to study AAV vector biology and develop novel AAV vector.
AAV DNA/RNA Barcode-Seq
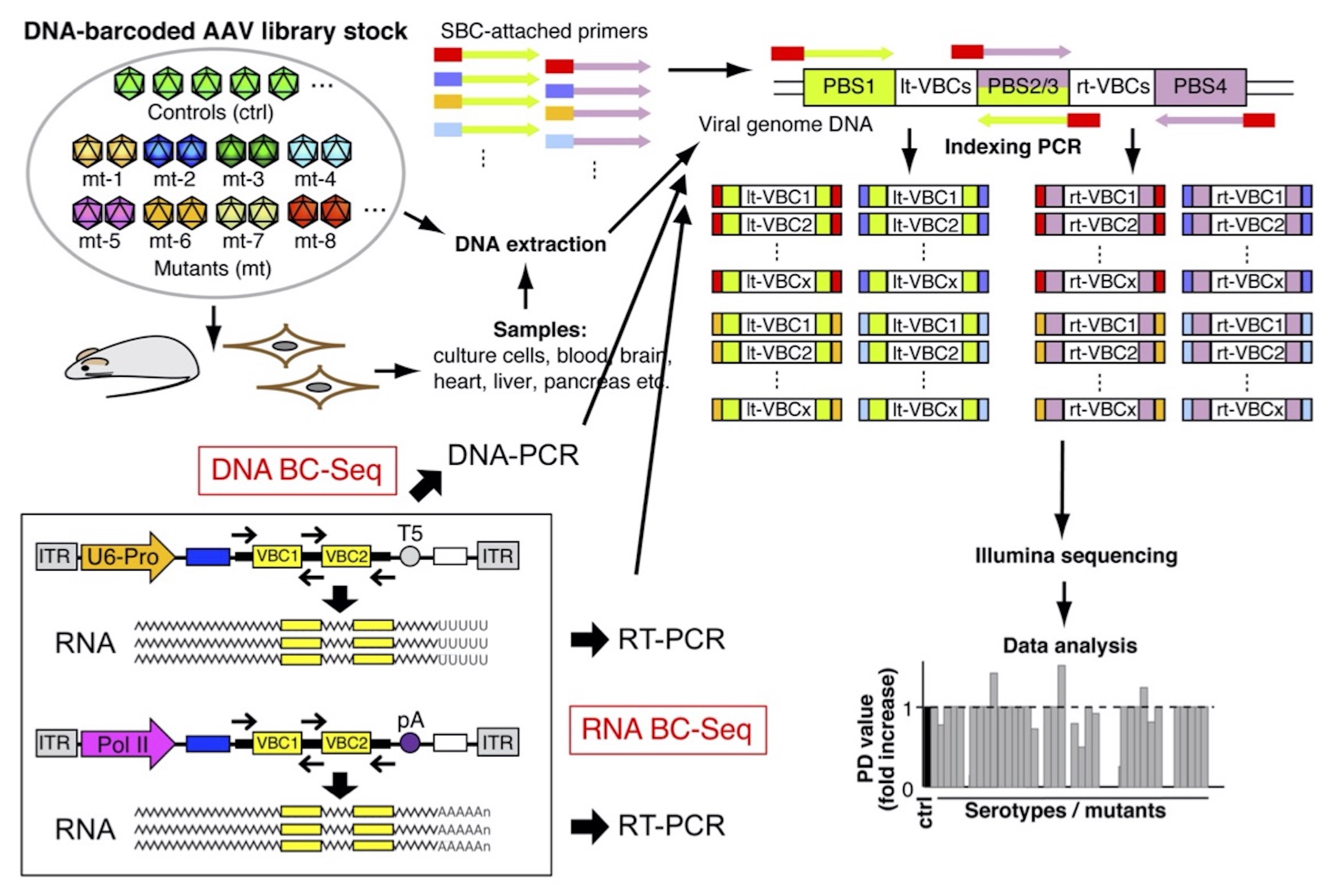
AAV Barcode-Seq. AAV Barcode-Seq is a next-generation sequencing (NGS)-based technologies that allows us to comprehensively determine genotype-phenotype relationships of AAV capsids derived from many serotypes, variants and mutants. This approach combines DNA barcode-tagged mutagenesis and multiplexed Illumina sequencing, making it possible to simultaneously characterize the multifaceted biological phenotypes of hundreds of AAV strains in various contexts, in a high-throughput manner, using a small number of replicates. In this approach, an AAV library is produced that comprises many different AAV strains whose viral genome contain strain-specific DNA barcodes that are transcribed either a ubiquitous RNA Pol III promoter (e.g., the human U6 snRNA promoter) or a ubiquitous (e.g., the CAG promoter) or cell-type specific RNA Pol II promoter (e.g., the human synapsin I promoter). Our AAV Barcode-Seq system uses a pair of left (lt) and right (rt) viral clone-specific 12 nucleotide-long DNA barcodes (Virus Bar Code or VBC) that are transcribed into RNA barcodes. Sample barcodes (SBCs) allows sample multiplexing. The principle of AAV Barcode-Seq is a bioinformatic comparison of the input library to the AAV capsid sequences recovered as viral genome DNA or as viral genome transcripts from the experimental samples. Using the same principle as RNA-Seq uses, AAV Barcode-Seq quantifies phenotypic differences (PDs) between different AAV strains as a function of strain demographics.
Adachi, K., Enoki, T., Kawano, Y., Veraz, M. G., Nakai, H. (2014) Drawing a high resolution functional map of adeno-associated virus capsid by massively parallel sequencing. Nat. Commun. 5:3075.
Adachi, K., Ido, H., Holman, T.W., Nakai, H. Development of a universal AAV Barcode-Seq system expressing RNA barcodes for AAV vector research. Oregon Health and Science University Research Week, Portland, OR, May 5-9, 2014.
Huang, S. Adachi, K., Baggett, H. R., Song, Z., Dissen, D. A., Ojeda, S. R., Nakai, H. Cell Type-Specific TRAnscription-Dependent Directed Evolution (TRADE) Identifies Novel AAV Capsids Capable of Enhanced Neuronal Transduction in Mice and Non-Human Primates. The 22th Annual Meeting of the American Society of Gene and Cell Therapy, Washington D.C., April 29 - May 2, 2019.
Capsid functional mapping
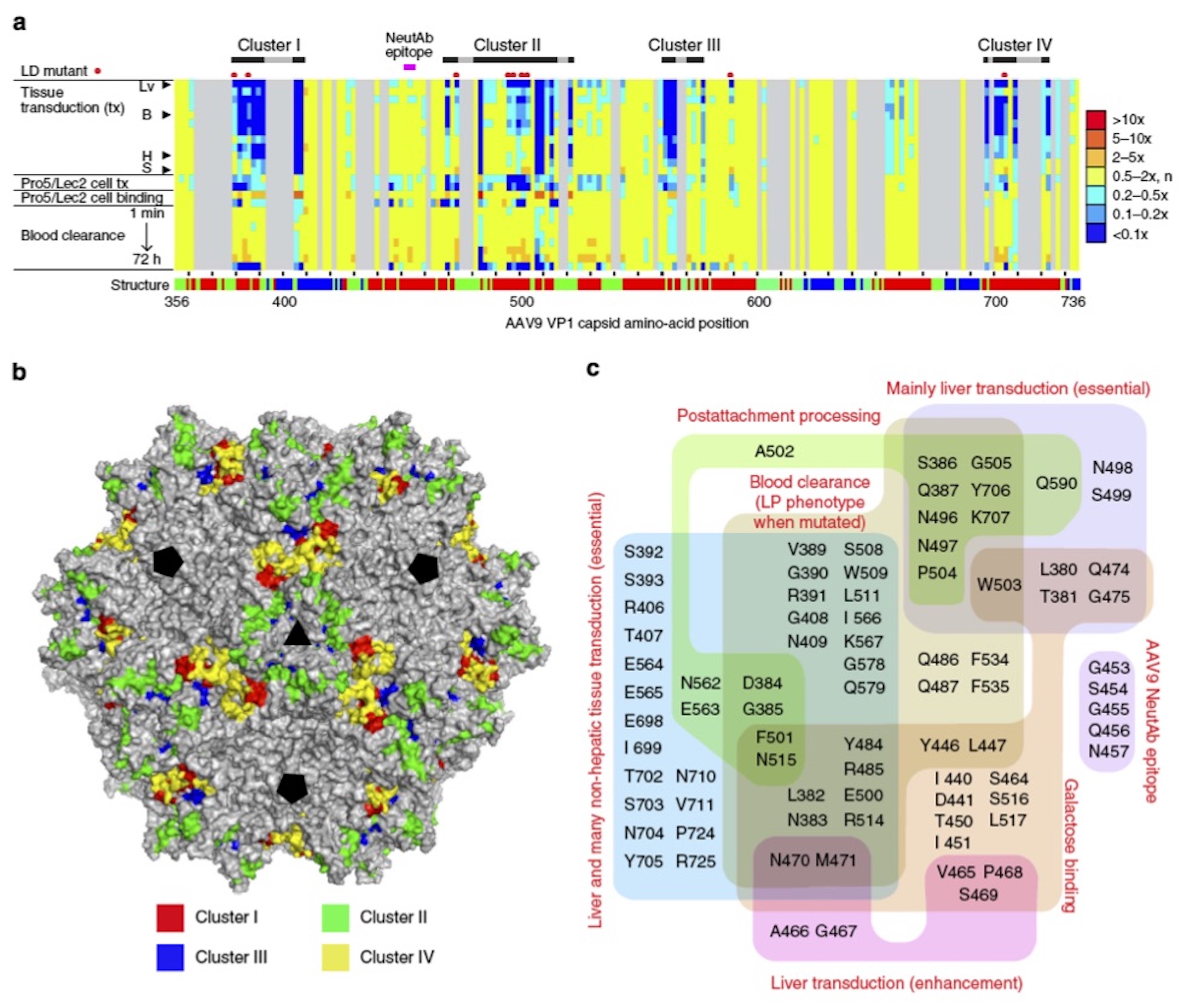
Functional maps of the C-terminal half of the AAV9 capsid amino acids. (a) A 2D heat map showing correlations between positions of AA mutations and phenotypic changes. Functionally important amino acids form four clusters (Clusters I–IV). Lv, liver; B, brain; H, heart; S, spleen. (b) A 3D map on the full AAV9 capsid atomic model showing topological locations of the functionally important amino acids in Clusters I–IV. (c) A Venn diagram showing correlation between AAV9 capsid amino-acid residues and various AAV9 phenotypes.
Adachi, K., Enoki, T., Kawano, Y., Veraz, M. G., Nakai, H. (2014) Drawing a high resolution functional map of adeno-associated virus capsid by massively parallel sequencing. Nat. Commun. 5:3075.
Phenotypic clustering of AAV capsid mutants
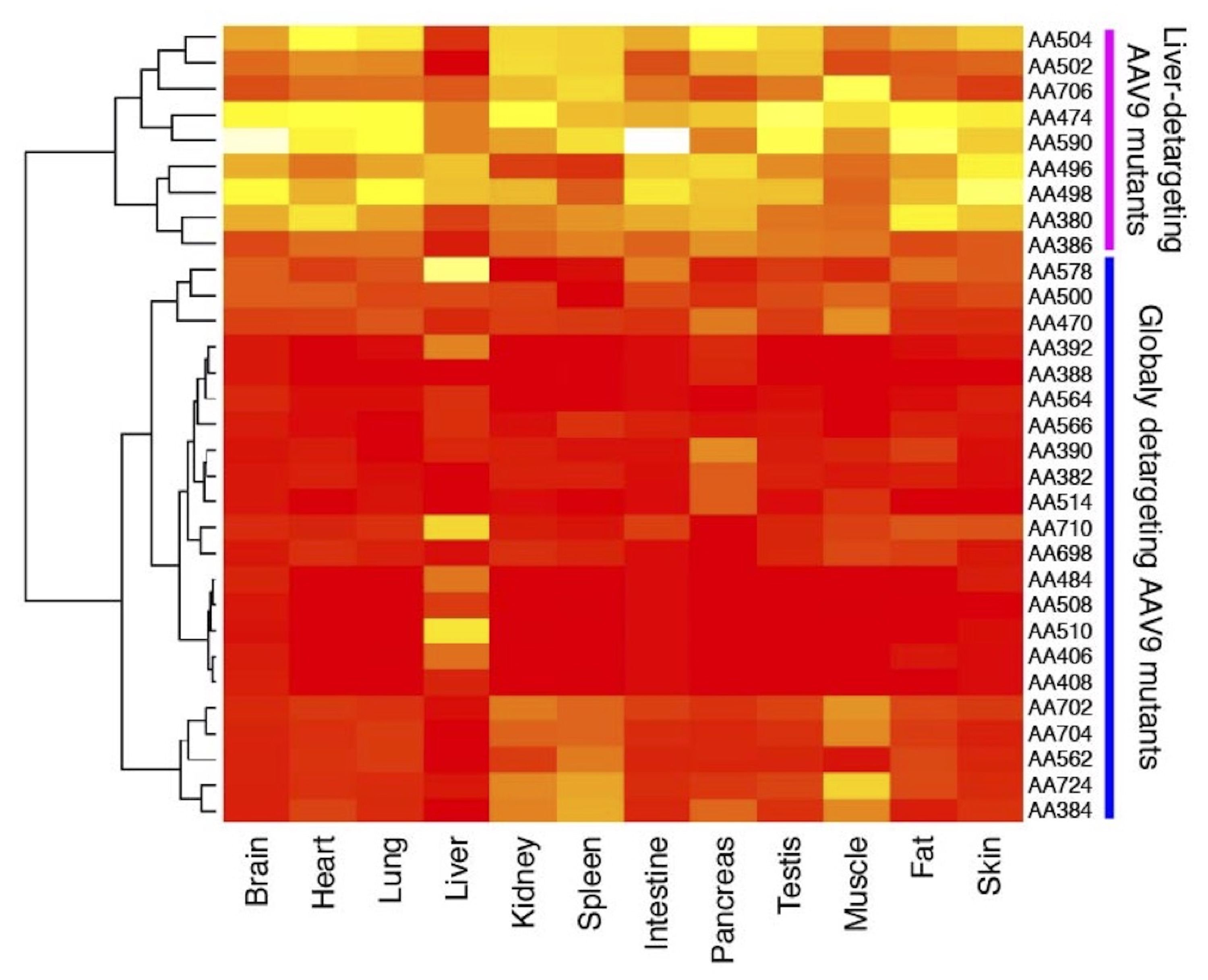
Hierarchical clustering analysis of liver-detargeted AAV9 double alanine (AA) mutants. The mutants can be grouped into two phenotypically distinct groups: liver-detargeting (LD) mutants and globally detargeting (GD) mutants. The LD mutants mainly detarget the liver while GD mutants show impaired transduction in many tissues in addition to the liver.
Adachi, K., Enoki, T., Kawano, Y., Veraz, M. G., Nakai, H. (2014) Drawing a high resolution functional map of adeno-associated virus capsid by massively parallel sequencing. Nat. Commun. 5:3075
Molecular imaging of AAV transduction by AAV Barcode-Seq and NGS
Three-dimensional heat maps of AAV vector transduction in the brain of rhesus macaque following intra-cisterna magna injection of DNA-barcoded AAV library. AAV library-transduced brain hemisphere was cut into >100 small cubes and subjected to AAV Barcode-Seq analysis. The Barcode-Seq data obtained from the cubes was reconstructed as a three-dimensional image and were superimposed on the brain MRI image.
Adachi, K., Liu Z., Kroenke C.D., Dissen, G.A., Ojeda, S.R., Nakai, H. Three-dimensional imaging of multiple AAV serotype/mutant vector transductions in the non-human primate brain by massively parallel sequencing. The 18th Annual Meeting of the American Society of Gene and Cell Therapy, New Orleans, LO, May 13-16, 2015. (Selected for presentation at the Presidential Symposium.)
A machine learning-based approach to understand pre-existing anti-AAV humoral immunity
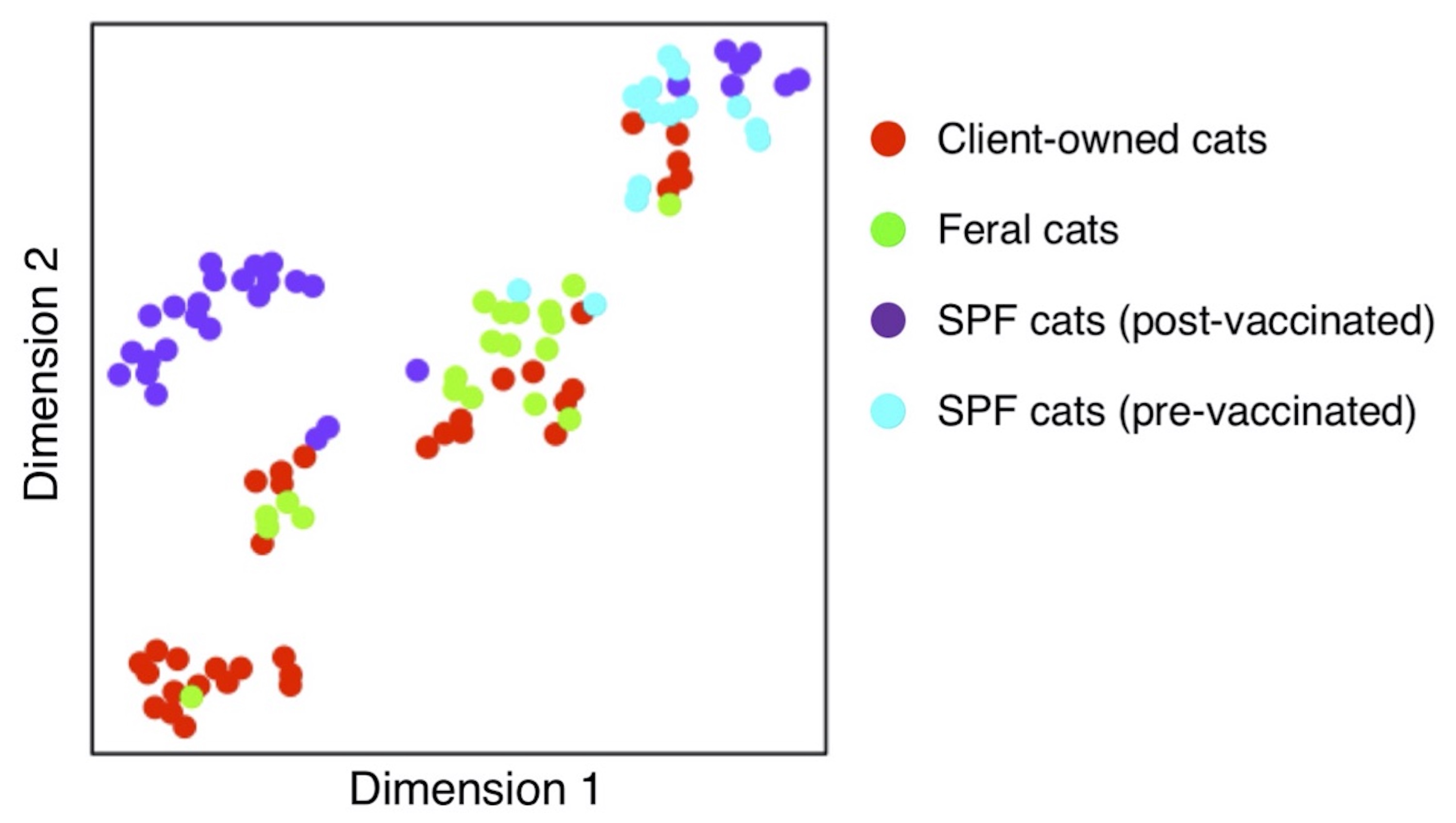
Five distinct cat groups identified based on humoral immunity profiles against multiple AAV serotypes. A comprehensive spectral dataset of serum levels of anti-AAV antibodies against AAV serotypes 1 to 11 in four different cat populations were subjected to dimensionality reduction using t-distributed stochastic neighbor embedding machine learning algorithm. Five distinct groups were revealed, providing insights that help understand pre-existing anti-AAV humoral immunity in the four cat populations that are different in nature.
Adachi, K., Dissen, G.A., Lomniczi, A., Xie, Q., Ojeda, S.R., Nakai, H. (2020) Adeno-associated virus-binding antibodies detected in cats living in the Northeastern United States lack neutralizing activity. Sci. Rep. (accepted).
AAV Vector Development
AAV Vector Development
AAV vectors made of naturally occurring AAV serotypes have been used in human gene therapy clinical trials and three AAV vector products have been commercialized (AAV1-Glybera, discontinued; AAV2-Luxturna; AAV9-Zolgensma) by 2019. In addition, ongoing preclinical studies of AAV vector-mediated gene therapy using animal models of human diseases continually report exciting outcomes. Thus, AAV vectors have gained increasing attention as a promising gene delivery vehicle for human gene therapy. However, various issues to overcome still remain in order to broaden the success of AAV-based gene therapy. These issues include:
- The presence of many extracellular and intracellular barriers (physical and biological) that hinder efficient gene delivery to target cells/tissues, necessitating administration of high vector doses for clinically beneficial outcomes
- Substantial vector spillover to non-target cells/tissues at therapeutically effective vector doses due to promiscuous viral tropism
- Efficacy-limiting host immune responses against viral proteins
- The high prevalence of preexisting anti-AAV neutralizing antibodies in humans
- A potential risk of AAV vector-mediated insertional mutagenesis causing malignancy
- Cost-effective high titer AAV vector production is challenging
To tackle these issues, we take multi-disciplinary experimental and analytical approaches using high-throughput molecular, cellular, and structural biology techniques coupled with bioinformatics and computational science.
Due to the simplicity of structures, AAV vectors can be engineered easily. In addition, studies have demonstrated that even a very small change in the capsid amino acid sequence, even a single amino acid mutation, can drastically change biological phenotypes. Thus, the Nakai lab mainly focuses on genetic engineering of AAV vector components to overcome these issues.
Specific areas of interest
Knowledge-based capsid design to modify AAV vector tropism
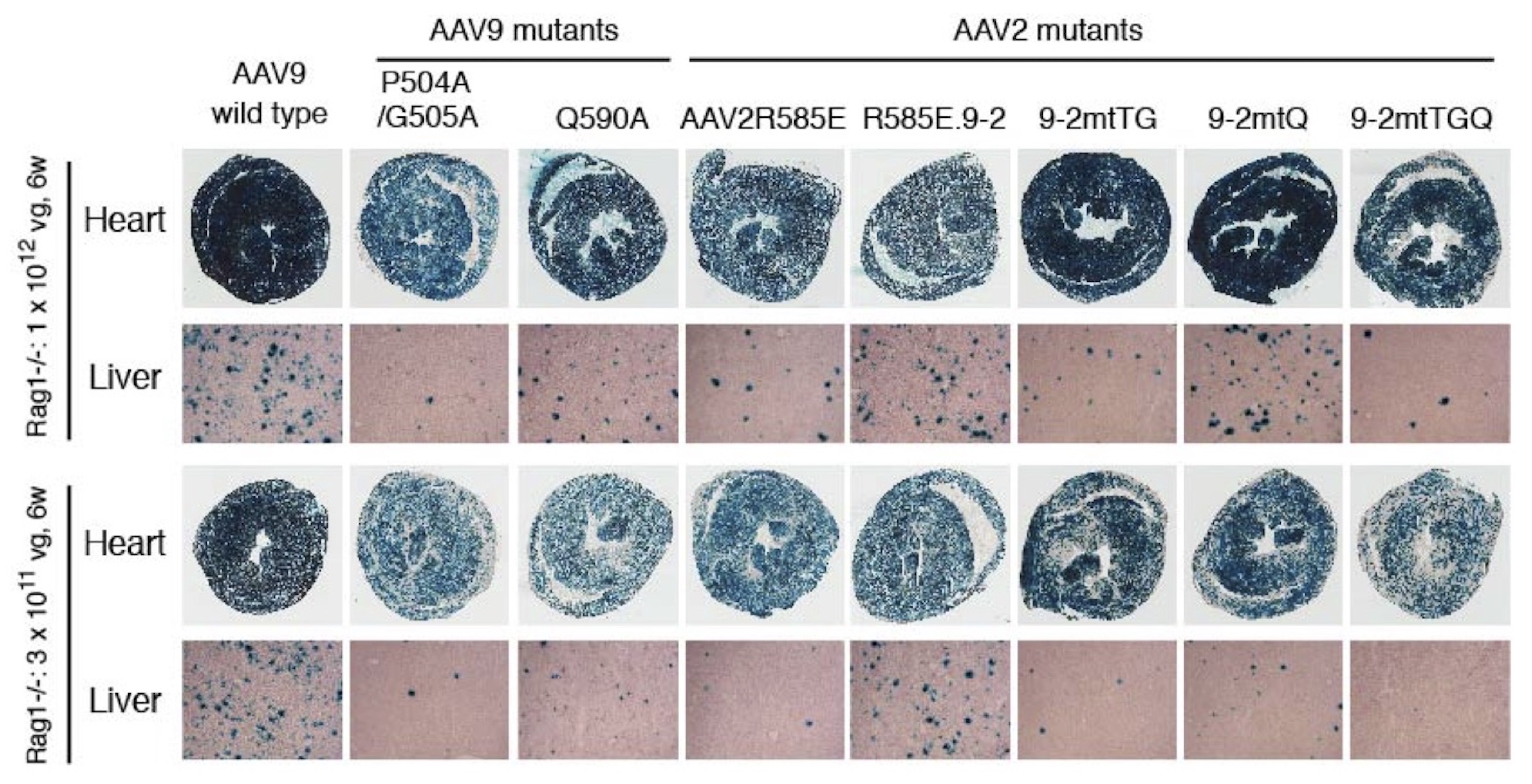
Liver and heart transduction with AAV9 and AAV2R585E mutants. Rag1 -/- mice were injected with AAV-CMV-lacZ vector packaged with various AAV capsids at a dose of 3x1011 or 1x1012 vg per mouse intravenously. Tissue transduction was assessed by X-Gal staining. AAV2R585E.9-2 is a heparin binding-defective AAV2 mutant carrying a galactose binding motif. The amino acid positions composing a galactose binding motif could be identified by AAV Barcode-Seq. Liver-detargeting amino acid changes identified in the AAV9 capsid by AAV Barcode-Seq (P504A, G505A and Q590A) were introduced in AAV2R585E.9-2 (TG, Q and TGQ mutants), which could substantially reduce liver transduction while preserving the cardiotropic property of the capsids.
Adachi, K., Enoki, T., Kawano, Y., Veraz, M. G., Nakai, H. (2014) Drawing a high-resolution functional map of adeno-associated virus capsid by massively parallel sequencing. Nat. Commun. 5:3075.
TRAnscription-dependent Directed Evolution (TRADE)
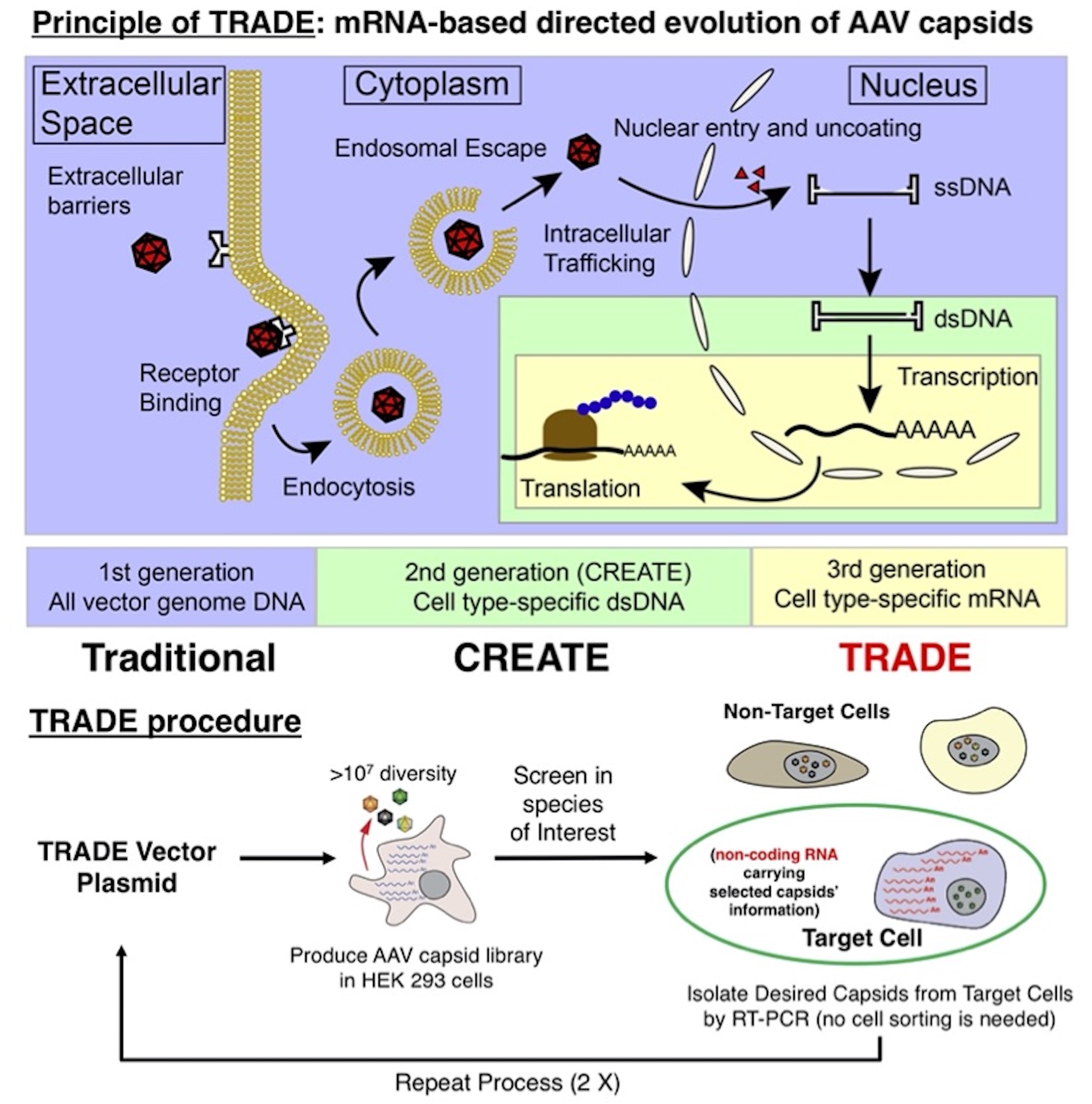
Directed evolution of the AAV capsid is a powerful tool for identifying AAV variants with enhanced transduction; however, its application in large animals, including non-human primates, has remained a challenge. In order to overcome this limitation, we developed a next-generation directed evolution system that selects enhanced capsids at the level of vector-mediated cell type-specific transgene expression in any animal species. TRADE does not require the use of a recombinase to confer cell-type specificity; therefore TRADE significantly improves on currently available AAV capsid directed evolution approaches, including the recently described CREATE system. The TRADE system utilizes a bicistronic vector genome containing (1) an AAV2 p40-driven cap open reading frame, and (2) a cell type-specific enhancer and promoter drives expression of the capsid VP protein sequence as a non-coding RNA. This vector configuration allows the recovery of capsid sequences from AAV variants that are capable of mediating cell type-specific mRNA expression by RT-PCR, and minimizes expression of immunogenic AAV capsids. In the proof-of-concept study, we sought to identify novel AAV capsids with enhanced neuronal transduction by utilizing a human synapsin I (hSynI) promoter in the TRADE system. We generated an AAV hSynI TRADE library consisting of an AAV9 liver-detargeted platform with an 8-amino acid peptide display inserted at position Q588 and flanked by glycine-serine linkers. We injected C57BL/6 mice and one rhesus macaque intravenously with this AAV library, harvested brain tissues 12 days post-injection, and recovered AAV capsid sequences from transduced hSynI-expressing brain neurons by RT-PCR. As a result, we have so far identified a novel mutant, AAV-HN1, that transduces brain neurons in C57BL/6 mice, BALB/c mice and rhesus macaque substantially better than AAV9 with much higher neuron specificity, following IV injection. Because the TRADE system does not require recombinase, it can be readily applied to a variety of animal species and cell types to rapidly advance the development of enhanced AAV vectors for research and clinical applications. Presented by Samuel Huang at the ASGCT2019 and 2020 Annual Meetings.
High-throughput mapping of anti-AAV capsid polyclonal antibody conformational epitopes by NGS (IP-Seq)
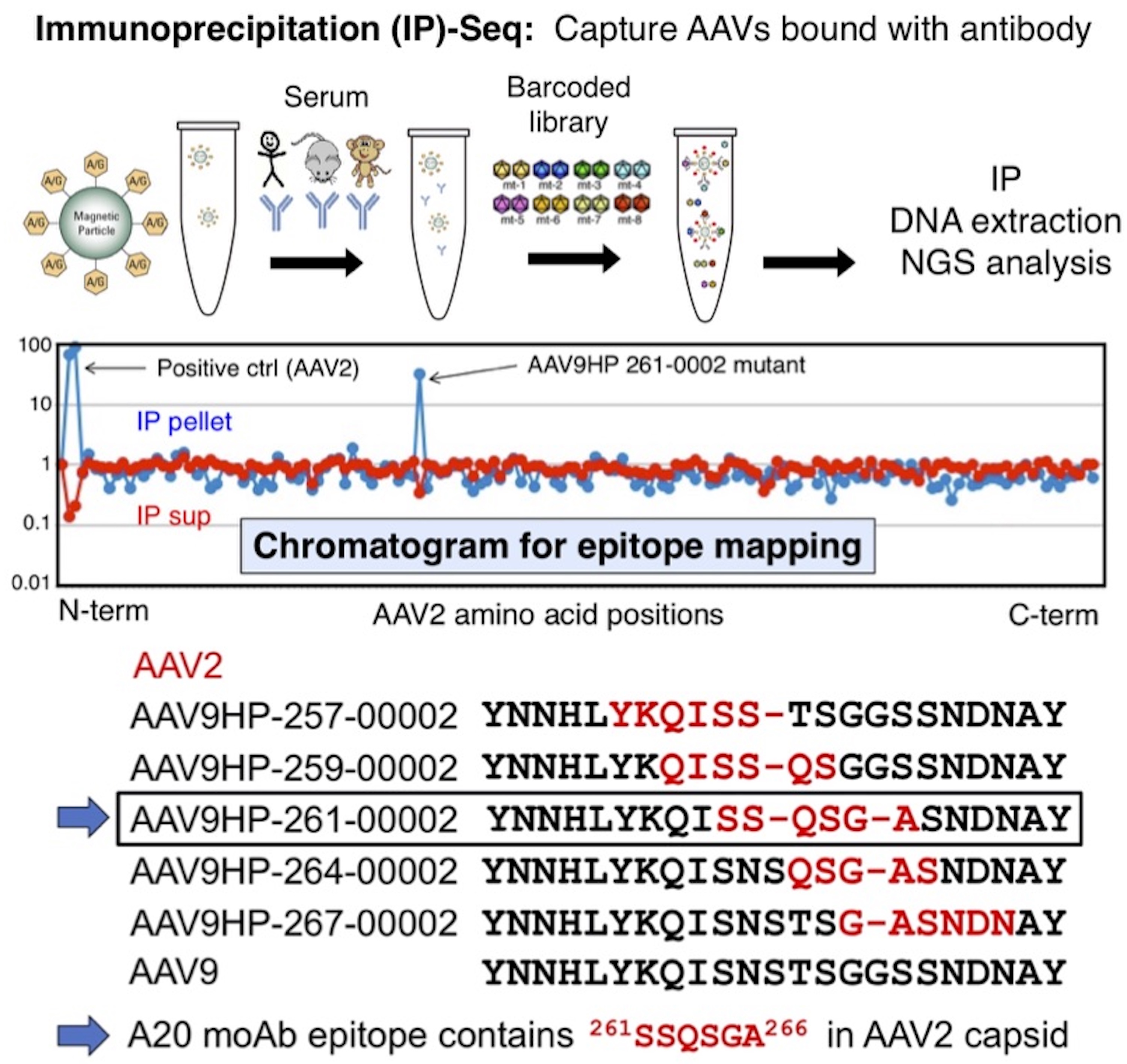
A proof-of-concept of IP-Seq experiment using a commercially available mouse monoclonal antibody against AAV2 capsid, A20. Epitopes for polyclonal anti-AAV neutralization antibodies can be identified by epitopes can be identified by Pharmacokinetic (PK)-Seq, an in vivo functional assay using mice that employs the same principle as that for IP-Seq. Comprehensive information about epitopes for polyclonal anti-AAV antibodies will be utilized to design novel AAV capsids that can escape neutralization (i.e., stealth AAV capsids).
Chang X.L., Adachi, K., Keck, I., Nakai, H. Mapping of conformational epitopes of monoclonal and polyclonal antibodies against AAV capsids by an Immunoprecipitation-Seq (IP-Seq) technology. The 20th Annual Meeting of the American Society of Gene and Cell Therapy, Washington D.C., May 10-13, 2017.
Baggett, H. R., Chang, X. L., Adachi, K., Nakai, H. Human polyclonal anti-AAV neutralizing antibody epitope mapping by NGS identifies common epitopes and enables the design of stealth mutants. The 22th Annual Meeting of the American Society of Gene and Cell Therapy, Washington D.C., April 29 - May 2, 2019.
Gene Therapy
Gene Therapy
In addition to research on AAV biology and novel gene therapy development, the Nakai lab has multiple disease-focused projects aiming at developing novel AAV vector-mediated gene therapy for various human diseases in collaboration with experts in each field. The projects have been supported by both industry-sponsored grants, NIH, and non-profit funding agencies. Target diseases include: CNS diseases including lysosomal storage diseases, Rett syndrome, CDKL5 deficiency, mitochondrial encephalomyopathy, metabolic liver diseases, retinal degeneration, type I diabetes, chronic pain, chronic kidney diseases. The Nakai lab is extremely enthusiastic about exploring new collaboration opportunities with disease experts in various fields.
Specific areas of interest
Gene therapy for type I diabetes
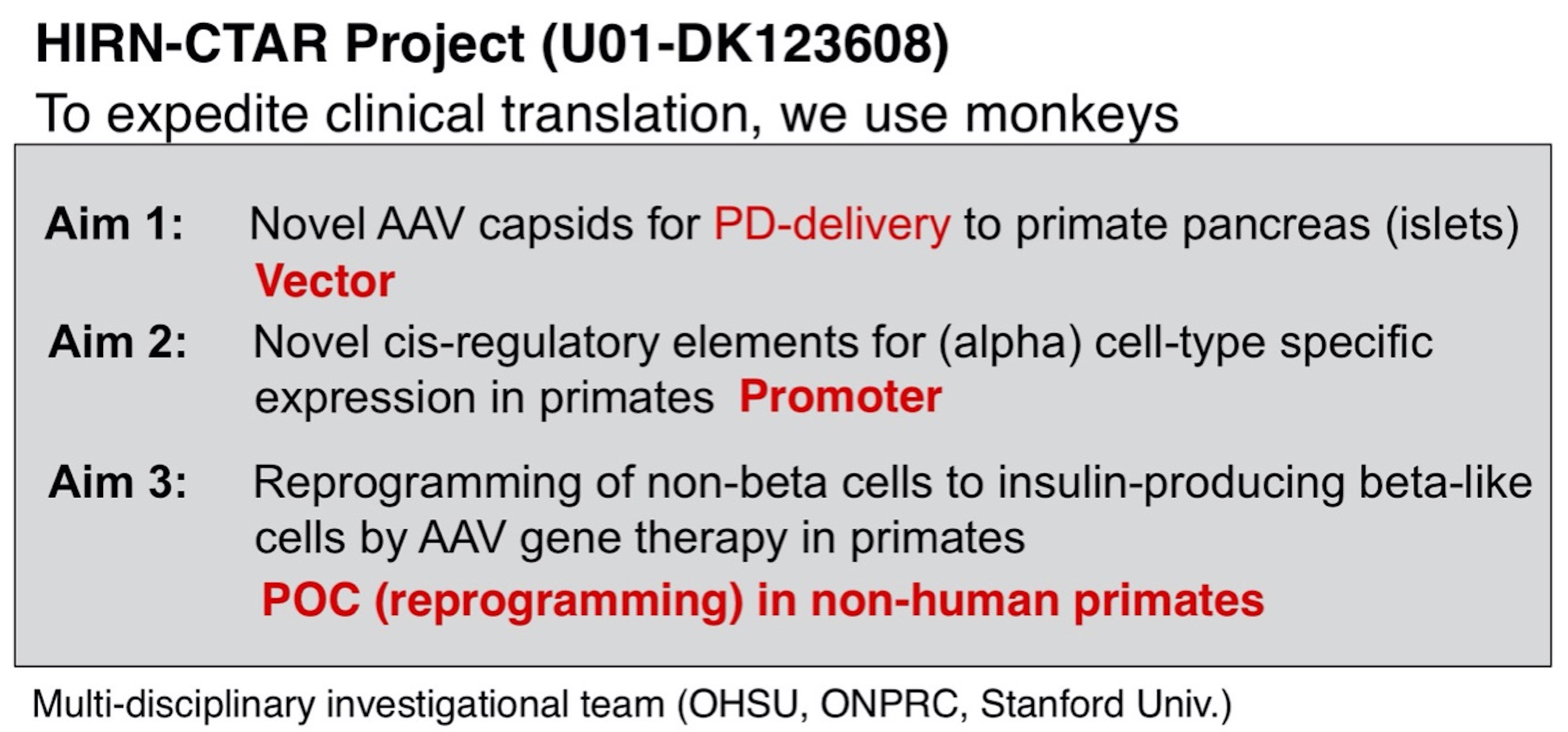
Type 1 diabetes (T1D) is a chronic incurable disease that requires daily insulin injection and blood sugar management. Even with proper management the disease often causes complications, leading to more premature mortality than in the general population. Remarkable progress has been made recently in understanding of pancreatic islet cell development and regeneration at the molecular level, which has enabled the creation of insulin-producing β-like cells from non-β-cells by viral vector-mediated delivery of key transcription factors. This approach is capable of correcting diabetes in murine models. Together, these findings provide a strong rationale for the development of gene therapy approaches for T1D. Adeno-associated virus (AAV) represents the most promising candidate for this purpose due to its clinically established safety and its superb ability to deliver genetic payloads to target cells. However, AAV vectors that deliver therapeutic molecules efficiently and specifically to the target cells in the pancreas (α, β, acinar, and duct cells) in primates including humans do not currently exist. In order to significantly expedite clinical translation of AAV T1D gene therapy, the Nakai lab in collaboration with the Grompe lab (OHSU) and the Kay lab (Stanford), seeks to develop AAV vector approaches capable of delivering genetic payloads to, and mediating their expression in, the pancreatic target cells in humans efficiently, specifically and safely by retrograde pancreatic duct (PD) injection. This method can maximize local transduction and minimize vector dissemination. To achieve this goal, we will utilize non-human primates (NHPs) as well as human islets explants and humanized mice transplanted with human islets. Ultimately, we will show proof-of-concept of AAV vector-mediated approach to reprogram α-cells to β-like cells in NHPs. The investigational team belongs to the Human Islet Research Network, Consortium on Targeting and Regeneration (HIRN-CTAR).
AAV9-CAG-TdTomato vector injection via the pancreatic duct (PD) in a rhesus macaque. AAV9 predominantly transduces acinar cells following PD injection in rhesus macaques. We have identified AAV strains that transduce pancreatic islets in rhesus macaque following PD injection substantially better than AAV9 by AAV DNA/RNA Barcode-Seq
