Korkola Research
Dr. Korkola works on primarily on breast cancer, with a focus on how interactions with microenvironment impact the phenotype of cancer cells, emphasizing those that alter the way cells respond to therapy. Dr. Korkola’s lab also is interested in understanding the mechanisms of resistance induced by extrinsic and intrinsic factors and developing multidrug combinations to overcome resistance and improve patient outcomes.
Cancer Research United Kingdom Project

In September 2018, Dr. Korkola won a new grant jointly awarded to himself and Daniel Muñoz-Espín, Ph.D., Programme Group Leader, at the Cancer Research UK Early Detection Programme. The proposed research will used microenviroment microarrays (MEMA) technology to examine causes of cancer based on the microenvironment, specifically certain cell secretions called senescence-associated secretory phenotype (SASP) factors. MEMA technologies the Korkola Lab has developed will allow the researchers to screen thousands of unique combinations of matrix proteins and soluble ligands for their impact on cells.
MEMA will be used to identify SASP factors that can drive the production of more cancer cells (proliferation), the development of non-specific cells into cancerous cell (differentiation changes), and increased invasiveness in precancerous and early lung and breast cancer cells. Three specifics aims:
- Define alterations in the early pre-neoplastic microenvironment that could be used to screen for individuals at high risk for the formation of cancerous lesions.
- Define factors in the early pre-neoplastic microenvironment that could be targeted for cancer prevention and diagnosis.
- Identify factors and related signalling pathways that are important in driving the progression of pre-neoplastic lesions into fully invasive neoplasms, allowing potential targeting of these factors and/or pathways for therapeutic interventions.
Overall, these studies will enable us to identify new causes of early tumor formation (tumorigenesis), provide new markers for detection of early breast and lung lesions, and provide new drug targets for preventing tumors.
Read about the CRUK-OHSU Project Award granted to Dr. Korkola
Read more about CRUK Early Detection
Microenvironment and cancer
Microenvironment Microarrays (MEMA)
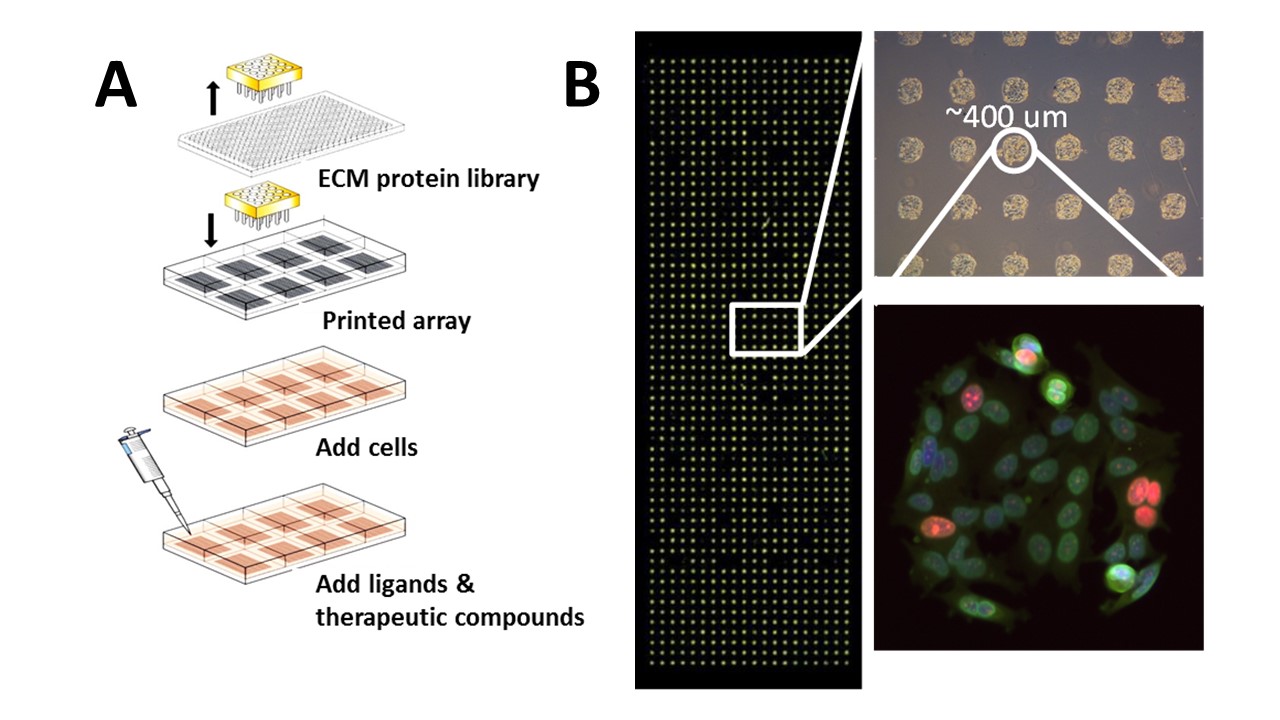
The Korkola Lab has spent the past several years developing microenvironment microarray, or MEMA technology. This approach (see Fig. 1 below for overview) allows us to construct thousands of simple combinatorial microenvironments on a single plate in which we can culture cells. We perform high content imaging on fixed and stained cells on MEMA to determine how the microenvironment alters the phenotypes of interest in both normal and cancer cells. We have used this approach to look at effects on proliferation, differentiation, spatial organization, and response to therapeutics in breast, prostate, pancreatic, and bladder cells.
HER2+ breast cancer
Much of the focus of the Korkola Lab over the past several years has been on HER2+ breast cancer (breast cancer that tests positive for human epidermal growth factor receptor 2, or HER2), which accounts for about 20 to 25% of all breast cancer cases and is associated with an aggressive phenotype, or form of cancer. Previously, we investigated the effects of mutations in the PI3K pathway (an intracellular signaling pathway important in regulating the cell cycle) on response.
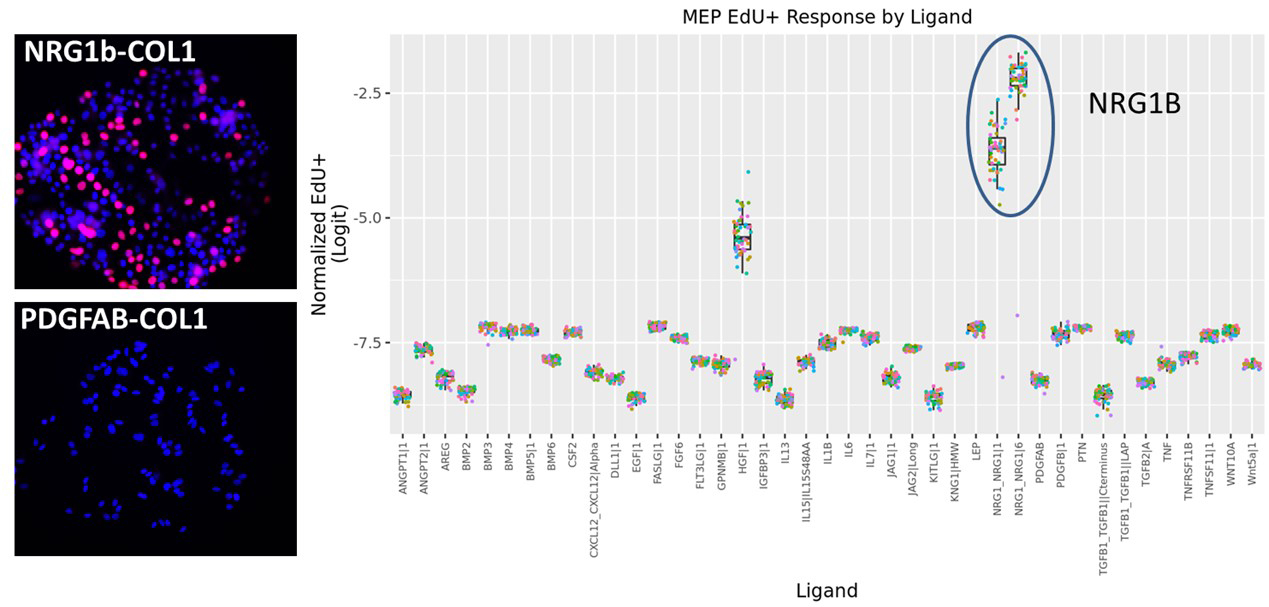
Currently, we are investigating differentially expressed genes associated with resistance, the effects of the microenvironment on drug response (Fig. 2 below), the effects of drug timing in combinatorial treatments, and novel drug treatments for resistant cancer.
Read OHSU News press release, "Area surrounding a tumor impacts how breast cancer cells grow"
Read OHSU blog post, "Study reveals factors outside of tumors that steer breast cancer aggressiveness"
Cancer heterogeneity
Measuring, Modeling and Controlling Heterogeneity - Cancer Systems Biology Center
Dr. Korkola is a co-principal investigator on the OHSU Measuring, Modeling and Controlling Heterogeneity (M2CH) Cancer Systems Biology Center (CSBC) project aimed at understanding how the microenvironment influences the heterogeneity and therapeutic response in triple negative breast cancer.
A triple negative breast cancer is any breast cancer that does not respond to medicines that work well on, or tests negative for the three common types of chemotherapy targets: estrogen receptors (making them ER negative), progesterone receptors (PR negative), or HER2 receptors (HER2 negative) These cancers, therefore, are harder to treat, since existing drugs that we have do not work against them. Triple negative breast cancers also tend to be more aggressive, or grow faster, than other breast cancers.