People and Labs
Collaborative from conception
The OCSSB was established in 2011 as a joint project between the Knight Cancer Institute and the School of Medicine. Thus, while it is part of the Department of Biomedical Engineering, the OCSSB maintains strong relationships with other units within OHSU, particularly the Knight Cancer Institute and the Computational Biology program. Within the Knight Cancer Institute, the OCSSB collaborates with the Cancer Early Detection Advanced Research (CEDAR) center and Precision Oncology program for many projects.
View OCCSB people, including:
Kimberly Beatty Lab

Develop chemical probes that advance our understanding of human diseases. Innovative approaches to molecular imaging of various targets, from classes of enzymes to multi-protein complexes. Learn more about the Beatty Lab.
Young Hwan Chang Group

Computational and interdisciplinary approaches to studying cancer. One main focus is to develop quantitative image analysis tools and build spatial systems biology models in order to understand how cancer cells adapt to their microenvironment (ME) and how tumor cell-ME interactions influence tumor cell physiology and response to therapy. Read more about the Chang Lab .
Catherine Galbraith Laboratory

Molecular mechanisms of cellular decision processes, mechano-biology of disease, and development of advanced light microscopy techniques. Visit the Galbraith Lab website.
Jim Galbraith Laboratory

Combines engineering, biology, biophysics and optics to investigate fundamental mechanisms of processes such as wiring the brain, synaptogenesis, cell division, and cell motility in both health and disease at the single molecule level. Visit the Galbraith Lab website.
Summer Gibbs Laboratory

Development of optimized imaging reagents to expand the capabilities of macroscopic and microscopic cancer imaging. Fluorophore development for image-guided surgery, superresolution microscopy, and correlative light and electron microscopy to visualize and characterize cancer. Learn more about the Gibbs Lab.
Joe Gray Laboratory

Develop more durable and tolerable control of advanced breast and pancreatic cancers. Learn more about the Gray Lab.
Laura Heiser Laboratory
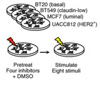
Identifying predictors of drug response and resistance, using novel imaging techniques to identify phenotypic changes associated with molecular aberrations and therapeutic response, and studying the influence of the microenvironment on cancer cells. Visit the Heiser Lab website.
Jim Korkola Laboratory

Breast cancer research with a focus on how interactions with microenvironment impact the phenotype of cancer cells, emphasizing those that alter the way cells respond to therapy. Learn more about the Korkola Lab.
Xiaolin Nan Laboratory
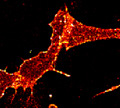
Single molecule spatial systems biology, with a particular interest in understanding how oncogenic signaling modules are assembled and operate in their cellular context and seeking their practical use in designing novel cancer therapeutics. Learn more about the Nan Lab.
Daniel Zuckerman Laboratory

Physics-based computational methods to study molecular, meso- and cell-scale systems. The group is particularly interested in the conformational behavior of proteins, in drug design for flexible receptors, as well as in molecular machines and their connections to cell behavior. Learn more about the Zuckerman Lab.